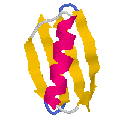
Protein G Background, References, and PDB Files
Protein G is a small globular protein produced by several Streptococcal species. The proteins are composed of two or three nearly identical domains of 55 amino acids each. The X-ray crystal and NMR structures displayed here are of the B1 domain from one of those species.
Protein G binds the Fc regions of IgG very tightly (Kd = 10-10 M); in this functional characteristic, they resemble the more famous Staphylococcal Protein A ("StaphA Protein") used as a molecular biology reagent. However, there is no sequence or structural similarity between Protein G and the all a-helix Protein A.
Structure Determination.
Gallagher, T. et al. (1994) "Two crystal structures of the B1 immunoglobulin-binding domain of Streptococcal protein G and comparison with NMR." Biochemistry 33, 4721-4729.
Gronenborn, A. M. et al. (1991) "A novel, highly stable fold of the immunoglobulin binding domain of Streptococcal protein G." Science 253 657.
Protein Folding Energetics.
Alexander, P. et al. (1992) "Thermodynamic analysis of the folding of the Streptococcal protein G IgG-binding domains B1 and B2: Why small proteins tend to have high denaturation temperatures." Biochemistry 31 3597.
Minor, D. L. & Kim, P. S. (1994) "Measurement of the b-sheet-forming propensities of amino acids." Nature 367 660.
Smith, C. K. et al. (1994) "A thermodynamic scale for the b-sheet forming tendencies of the amino acids." Biochemistry 33 5510.
Smith, C. K. & Regan, L. (1995) "Guidelines for protein design: The energetics of b-sheet side chain interactions." Science 270 980.
Minor, D. L. & Kim, P. S. (1996) "Context-dependent secondary structure formation of a designed protein sequence." Nature 380, 730.
The following Protein G (B1 domain) coordinate files (saved as MS Word (text) documents) were used for the various Protein G displays on these pages.
- 1pgb.pdb X-ray crystal structure from Gallagher, et al. (1994). Complete PDB file (pdb1pgb.ent); includes all 24 waters. The following are truncated versions of 1pgb.pdb:
- 2gb1.pdb NMR structure from Gronenborn, et al. (1991). Complete PDB file (pdb2gb1.ent); this is the average structure derived from the 60 calculated models and includes all protons on the protein (of course!) but no waters. The following are truncated versions of 2gb1.pdb:
Depending on how your Netscape preferences are set, a click on any of the above links will either (i) launch RasMol as a helper application, or (ii) fill your entire page with the structure using the Chime plugin.
An "option-click" on the above links will let you download and save the coordinates as a RasMol (RSML) file. You can then use RasMol on your computer to manipulate the the structure(s) in any way you choose.
Three extensions of this Protein Architecture tutorial are suggested:
- Beginner: Launch RasMol and open the 1pgb.pdb file with the first tutorial page in a separate Netscape window. Now use the RasMol Display and Color menus to reproduce the images shown on that tutorial page.
- Intermediate: Same drill as above, but now use statements from the RasMol command line to make images that are similar to those shown in the Secondary Structure or Tertiary Folding sections of the tutorial. Examples of some of the most-used commands are:
- select resno=23, select atomno=1, etc.
- restrict sheet, restrict helix, etc.
- set picking distance, set picking ident, etc.
- help topic (You specify the topic; RasMol provides explanations on the command line.)
More detailed help and explanation can be obtained as follows:
- Get a copy of the RasMol Manual: Frames or copy the "RasMol Manual" folder from the RasMac v2.6 folder on the BioServer.
- Open the manual with the "Open File..." option in the Netscape "File" menu.
- Open a PDB file with RasMac.
In this way you can quickly refer to the topics in the RasMol manual (displayed in the Netscape window), while you manipulate the PDB file (displayed in the RasMol windows).
- Advanced: Read Gallagher, et al. (1994) or Gronenborn, et al. (1991) and make displays of the many features of Protein G that were not shown in this tutorial.
If you want to compare the crystal and NMR structures in a side-by-side display using Rasmol, i.e. separate display windows, a simple trick is required: the RasMol program only allows one window to be open at a time. So, duplicate it and give the copy a different name e.g. "RasMac v2.6 copy". Now you can launch both programs at the same time and open different PDB files in each window.
 Return to Protein Architecture page.
Return to Protein Architecture page.