| |


- Files
- Control Panel
- Move & Rotate
- Display
- Rendering
- Tools
- Mutations
- Torsions
- Preferences
- hardware stereo
- electrostatics
- surface
- tips & tricks
Index







by N.Guex
& T.Schwede
|
|
Identifying Distorted Residues
When you are modelling a protein, or solving a structure,
it is always helpful to identify bad regions. Swiss-PdbViewer provides some
tools for that purpose.
Identifying Backbone Problems:
The best way to have a quick glance at the global problems is to color
the protein by "backbone problems". This will highlight disconnected regions
of the backbone, which is very useful during homology modelling, to help
you loacte and modify where the insrtions/deletions should be placed.
In addition, residues with a bad phi/psi conformation are also readily
identified; which might be useful during refinement of your structure.

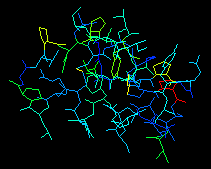
example of preliminary model analysis. Spots where insertions and
deletions will be needed appear in cyan; whereas residues with bad phi/psi
conformation appear in yellow (red for Prolines).
Identifying Distorted Residues.
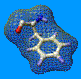
The best way to illustrate this is to give a practical example. Load
the protein 1CRN.pdb (which is included in the tutorial
package), and color it by Force Field Energy (but do only compute
bond and angles energies; and do not show the energy report). Overall,
the protein topology is correct, the residue with the highest energy (the
more bond distortion) beeing Proline 5.
Now Select the Arg17 only (which was blue, meaning that its bond length
and angles are quite good). Use the tool menu to "Shake the Selected Groups".
Apply a 0.2Å random displacement of any atom of Arg17. This means
that if you measure the RMS deviation between the unshaked residue and
the shaked residue, you will obtain a RMSd of 0.2. Indeed, by inspecting
your protein, it is hard to say that this Arg. is distorted.
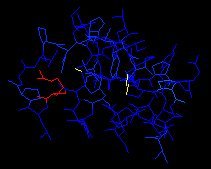
Now color your protein by force field energy. The distorted Arginie will
shine in bright red, whereas the rest of the protein is dark blue (except
Pro 5, which is blue).
|
 ExPASy Home page
ExPASy Home page ExPASy Home page
ExPASy Home page
 ExPASy Home page
ExPASy Home page