SWISS-PROT PROTEIN KNOWLEDGEBASE RELEASE 41.4 STATISTICS
1. INTRODUCTION
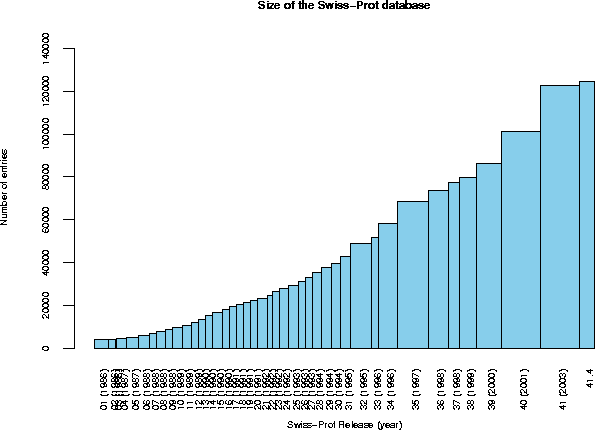
Release 41.4 of 11-Apr-2003 of Swiss-Prot contains 124464 sequence entries,
comprising 45704421 amino acids abstracted from 104368 references.
1905 sequences have been added since release 41, the sequence data of
178 existing entries has been updated and the annotations of
10818 entries have been revised. This represents an increase of 2%.
The growth of the database is summarized below.
 2. AMINO ACID COMPOSITION
2.1 Composition in percent for the complete database
Ala (A) 7.72 Gln (Q) 3.92 Leu (L) 9.57 Ser (S) 6.98
Arg (R) 5.24 Glu (E) 6.55 Lys (K) 5.97 Thr (T) 5.51
Asn (N) 4.28 Gly (G) 6.90 Met (M) 2.37 Trp (W) 1.18
Asp (D) 5.28 His (H) 2.26 Phe (F) 4.06 Tyr (Y) 3.12
Cys (C) 1.59 Ile (I) 5.89 Pro (P) 4.87 Val (V) 6.66
Asx (B) 0.000 Glx (Z) 0.000 Xaa (X) 0.01
2. AMINO ACID COMPOSITION
2.1 Composition in percent for the complete database
Ala (A) 7.72 Gln (Q) 3.92 Leu (L) 9.57 Ser (S) 6.98
Arg (R) 5.24 Glu (E) 6.55 Lys (K) 5.97 Thr (T) 5.51
Asn (N) 4.28 Gly (G) 6.90 Met (M) 2.37 Trp (W) 1.18
Asp (D) 5.28 His (H) 2.26 Phe (F) 4.06 Tyr (Y) 3.12
Cys (C) 1.59 Ile (I) 5.89 Pro (P) 4.87 Val (V) 6.66
Asx (B) 0.000 Glx (Z) 0.000 Xaa (X) 0.01
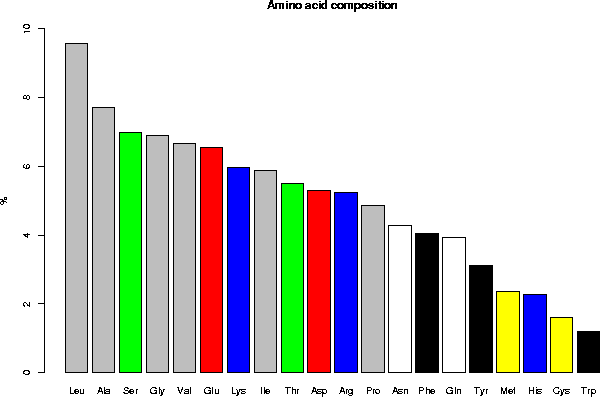 Legend: gray = aliphatic, red = acidic, green = small hydroxy,
blue = basic, black = aromatic, white = amide, yellow = sulfur
2.2 Classification of the amino acids by their frequency
Leu, Ala, Ser, Gly, Val, Glu, Lys, Ile, Thr, Asp, Arg, Pro, Asn, Phe,
Gln, Tyr, Met, His, Cys, Trp
3. TAXONOMIC ORIGIN
Total number of species represented in this release of Swiss-Prot: 7830
The first twenty species represent 52260 sequences: 42 % of the total
number of entries.
3.1 Table of the frequency of occurrence of species
Species represented 1x: 3701
2x: 1210
3x: 631
4x: 406
5x: 261
6x: 256
7x: 198
8x: 147
9x: 124
10x: 66
11- 20x: 330
21- 50x: 248
51-100x: 91
>100x: 161
3.2 Table of the most represented species
------ --------- --------------------------------------------
Number Frequency Species
------ --------- --------------------------------------------
1 9294 Homo sapiens (Human)
2 6247 Mus musculus (Mouse)
3 4893 Saccharomyces cerevisiae (Baker's yeast)
4 4832 Escherichia coli
5 3465 Rattus norvegicus (Rat)
6 2406 Bacillus subtilis
7 2321 Caenorhabditis elegans
8 2130 Schizosaccharomyces pombe (Fission yeast)
9 2092 Arabidopsis thaliana (Mouse-ear cress)
10 1794 Drosophila melanogaster (Fruit fly)
11 1773 Haemophilus influenzae
12 1530 Methanococcus jannaschii
13 1506 Escherichia coli O157:H7
14 1394 Bos taurus (Bovine)
15 1375 Mycobacterium tuberculosis
16 1257 Salmonella typhimurium
17 1064 Gallus gallus (Chicken)
18 1006 Escherichia coli O6
19 961 Shigella flexneri
20 920 Synechocystis sp. (strain PCC 6803)
21 880 Archaeoglobus fulgidus
22 846 Pseudomonas aeruginosa
23 845 Xenopus laevis (African clawed frog)
24 823 Sus scrofa (Pig)
25 796 Salmonella typhi
26 716 Aquifex aeolicus
27 705 Oryctolagus cuniculus (Rabbit)
28 687 Mycoplasma pneumoniae
29 681 Rhizobium meliloti (Sinorhizobium meliloti)
30 624 Vibrio cholerae
31 599 Treponema pallidum
32 586 Mycobacterium leprae
33 572 Buchnera aphidicola (subsp. Acyrthosiphon pisum)
34 560 Buchnera aphidicola (subsp. Schizaphis graminum)
35 544 Yersinia pestis
36 537 Helicobacter pylori (Campylobacter pylori)
37 535 Rickettsia prowazekii
38 526 Streptomyces coelicolor
39 520 Helicobacter pylori J99 (Campylobacter pylori J99)
40 498 Bacillus halodurans
41 495 Methanobacterium thermoautotrophicum
42 491 Zea mays (Maize)
43 489 Pasteurella multocida
44 486 Mycoplasma genitalium
45 469 Anabaena sp. (strain PCC 7120)
46 437 Lactococcus lactis (subsp. lactis) (Streptococcus lactis)
47 421 Thermotoga maritima
48 417 Oryza sativa (Rice)
49 406 Borrelia burgdorferi (Lyme disease spirochete)
50 405 Chlamydia trachomatis
51 403 Rhizobium sp. (strain NGR234)
52 394 Canis familiaris (Dog)
53 394 Neisseria meningitidis (serogroup B)
54 392 Chlamydia pneumoniae (Chlamydophila pneumoniae)
55 390 Neisseria meningitidis (serogroup A)
56 382 Chlamydia muridarum
57 370 Pyrococcus horikoshii
58 369 Caulobacter crescentus
59 367 Listeria monocytogenes
60 367 Clostridium acetobutylicum
61 363 Pyrococcus abyssi
62 363 Rhizobium loti (Mesorhizobium loti)
63 362 Ralstonia solanacearum (Pseudomonas solanacearum)
64 360 Listeria innocua
65 358 Streptococcus pneumoniae
66 356 Agrobacterium tumefaciens (strain C58 / ATCC 33970)
67 343 Nicotiana tabacum (Common tobacco)
68 342 Xylella fastidiosa
69 339 Deinococcus radiodurans
70 335 Xanthomonas campestris (pv. campestris)
71 333 Ovis aries (Sheep)
72 327 Staphylococcus aureus (strain N315)
73 326 Halobacterium sp. (strain NRC-1 / ATCC 700922 / JCM 11081)
74 326 Campylobacter jejuni
75 324 Staphylococcus aureus (strain Mu50 / ATCC 700699)
76 319 Clostridium perfringens
77 316 Dictyostelium discoideum (Slime mold)
78 312 Corynebacterium glutamicum (Brevibacterium flavum)
79 306 Sulfolobus solfataricus
80 304 Staphylococcus aureus (strain MW2)
81 299 Xanthomonas axonopodis (pv. citri)
82 291 Streptococcus pyogenes
83 289 Aeropyrum pernix
84 289 Pisum sativum (Garden pea)
85 285 Pyrococcus furiosus
86 279 Staphylococcus aureus
87 276 Brucella melitensis
88 268 Bacteriophage T4
89 266 Neurospora crassa
90 265 Triticum aestivum (Wheat)
91 264 Rickettsia conorii
92 264 Candida albicans (Yeast)
93 263 Thermoanaerobacter tengcongensis
94 258 Hordeum vulgare (Barley)
95 254 Vaccinia virus (strain Copenhagen)
96 254 Methanosarcina mazei (Methanosarcina frisia)
97 254 Glycine max (Soybean)
98 253 Methanosarcina acetivorans
99 251 Lycopersicon esculentum (Tomato)
100 248 Rhodobacter capsulatus (Rhodopseudomonas capsulata)
3.3 Taxonomic distribution of the sequences
Legend: gray = aliphatic, red = acidic, green = small hydroxy,
blue = basic, black = aromatic, white = amide, yellow = sulfur
2.2 Classification of the amino acids by their frequency
Leu, Ala, Ser, Gly, Val, Glu, Lys, Ile, Thr, Asp, Arg, Pro, Asn, Phe,
Gln, Tyr, Met, His, Cys, Trp
3. TAXONOMIC ORIGIN
Total number of species represented in this release of Swiss-Prot: 7830
The first twenty species represent 52260 sequences: 42 % of the total
number of entries.
3.1 Table of the frequency of occurrence of species
Species represented 1x: 3701
2x: 1210
3x: 631
4x: 406
5x: 261
6x: 256
7x: 198
8x: 147
9x: 124
10x: 66
11- 20x: 330
21- 50x: 248
51-100x: 91
>100x: 161
3.2 Table of the most represented species
------ --------- --------------------------------------------
Number Frequency Species
------ --------- --------------------------------------------
1 9294 Homo sapiens (Human)
2 6247 Mus musculus (Mouse)
3 4893 Saccharomyces cerevisiae (Baker's yeast)
4 4832 Escherichia coli
5 3465 Rattus norvegicus (Rat)
6 2406 Bacillus subtilis
7 2321 Caenorhabditis elegans
8 2130 Schizosaccharomyces pombe (Fission yeast)
9 2092 Arabidopsis thaliana (Mouse-ear cress)
10 1794 Drosophila melanogaster (Fruit fly)
11 1773 Haemophilus influenzae
12 1530 Methanococcus jannaschii
13 1506 Escherichia coli O157:H7
14 1394 Bos taurus (Bovine)
15 1375 Mycobacterium tuberculosis
16 1257 Salmonella typhimurium
17 1064 Gallus gallus (Chicken)
18 1006 Escherichia coli O6
19 961 Shigella flexneri
20 920 Synechocystis sp. (strain PCC 6803)
21 880 Archaeoglobus fulgidus
22 846 Pseudomonas aeruginosa
23 845 Xenopus laevis (African clawed frog)
24 823 Sus scrofa (Pig)
25 796 Salmonella typhi
26 716 Aquifex aeolicus
27 705 Oryctolagus cuniculus (Rabbit)
28 687 Mycoplasma pneumoniae
29 681 Rhizobium meliloti (Sinorhizobium meliloti)
30 624 Vibrio cholerae
31 599 Treponema pallidum
32 586 Mycobacterium leprae
33 572 Buchnera aphidicola (subsp. Acyrthosiphon pisum)
34 560 Buchnera aphidicola (subsp. Schizaphis graminum)
35 544 Yersinia pestis
36 537 Helicobacter pylori (Campylobacter pylori)
37 535 Rickettsia prowazekii
38 526 Streptomyces coelicolor
39 520 Helicobacter pylori J99 (Campylobacter pylori J99)
40 498 Bacillus halodurans
41 495 Methanobacterium thermoautotrophicum
42 491 Zea mays (Maize)
43 489 Pasteurella multocida
44 486 Mycoplasma genitalium
45 469 Anabaena sp. (strain PCC 7120)
46 437 Lactococcus lactis (subsp. lactis) (Streptococcus lactis)
47 421 Thermotoga maritima
48 417 Oryza sativa (Rice)
49 406 Borrelia burgdorferi (Lyme disease spirochete)
50 405 Chlamydia trachomatis
51 403 Rhizobium sp. (strain NGR234)
52 394 Canis familiaris (Dog)
53 394 Neisseria meningitidis (serogroup B)
54 392 Chlamydia pneumoniae (Chlamydophila pneumoniae)
55 390 Neisseria meningitidis (serogroup A)
56 382 Chlamydia muridarum
57 370 Pyrococcus horikoshii
58 369 Caulobacter crescentus
59 367 Listeria monocytogenes
60 367 Clostridium acetobutylicum
61 363 Pyrococcus abyssi
62 363 Rhizobium loti (Mesorhizobium loti)
63 362 Ralstonia solanacearum (Pseudomonas solanacearum)
64 360 Listeria innocua
65 358 Streptococcus pneumoniae
66 356 Agrobacterium tumefaciens (strain C58 / ATCC 33970)
67 343 Nicotiana tabacum (Common tobacco)
68 342 Xylella fastidiosa
69 339 Deinococcus radiodurans
70 335 Xanthomonas campestris (pv. campestris)
71 333 Ovis aries (Sheep)
72 327 Staphylococcus aureus (strain N315)
73 326 Halobacterium sp. (strain NRC-1 / ATCC 700922 / JCM 11081)
74 326 Campylobacter jejuni
75 324 Staphylococcus aureus (strain Mu50 / ATCC 700699)
76 319 Clostridium perfringens
77 316 Dictyostelium discoideum (Slime mold)
78 312 Corynebacterium glutamicum (Brevibacterium flavum)
79 306 Sulfolobus solfataricus
80 304 Staphylococcus aureus (strain MW2)
81 299 Xanthomonas axonopodis (pv. citri)
82 291 Streptococcus pyogenes
83 289 Aeropyrum pernix
84 289 Pisum sativum (Garden pea)
85 285 Pyrococcus furiosus
86 279 Staphylococcus aureus
87 276 Brucella melitensis
88 268 Bacteriophage T4
89 266 Neurospora crassa
90 265 Triticum aestivum (Wheat)
91 264 Rickettsia conorii
92 264 Candida albicans (Yeast)
93 263 Thermoanaerobacter tengcongensis
94 258 Hordeum vulgare (Barley)
95 254 Vaccinia virus (strain Copenhagen)
96 254 Methanosarcina mazei (Methanosarcina frisia)
97 254 Glycine max (Soybean)
98 253 Methanosarcina acetivorans
99 251 Lycopersicon esculentum (Tomato)
100 248 Rhodobacter capsulatus (Rhodopseudomonas capsulata)
3.3 Taxonomic distribution of the sequences
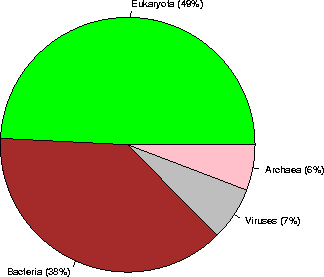 Kingdom sequences (% of the database)
Archaea 7218 ( 6%)
Bacteria 47533 ( 38%)
Eukaryota 61240 ( 49%)
Viruses 8473 ( 7%)
Within Eukaryota:
Kingdom sequences (% of the database)
Archaea 7218 ( 6%)
Bacteria 47533 ( 38%)
Eukaryota 61240 ( 49%)
Viruses 8473 ( 7%)
Within Eukaryota:
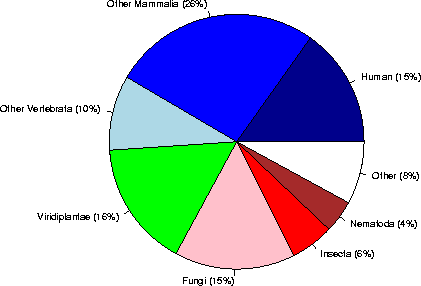 Category sequences (% of Eukaryota) (% of the complete database)
Human 9294 ( 15%) ( 7%)
Other Mammalia 16152 ( 26%) ( 13%)
Other Vertebrata 5824 ( 10%) ( 5%)
Viridiplantae 9814 ( 16%) ( 8%)
Fungi 9364 ( 15%) ( 8%)
Insecta 3394 ( 6%) ( 3%)
Nematoda 2535 ( 4%) ( 2%)
Other 4863 ( 8%) ( 4%)
3.4 Annotation of high-priority organisms
Category sequences (% of Eukaryota) (% of the complete database)
Human 9294 ( 15%) ( 7%)
Other Mammalia 16152 ( 26%) ( 13%)
Other Vertebrata 5824 ( 10%) ( 5%)
Viridiplantae 9814 ( 16%) ( 8%)
Fungi 9364 ( 15%) ( 8%)
Insecta 3394 ( 6%) ( 3%)
Nematoda 2535 ( 4%) ( 2%)
Other 4863 ( 8%) ( 4%)
3.4 Annotation of high-priority organisms
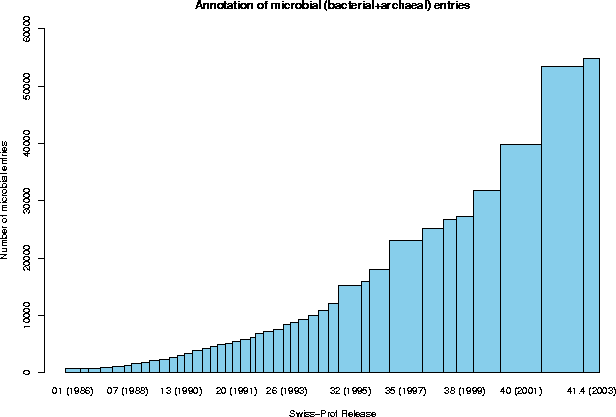
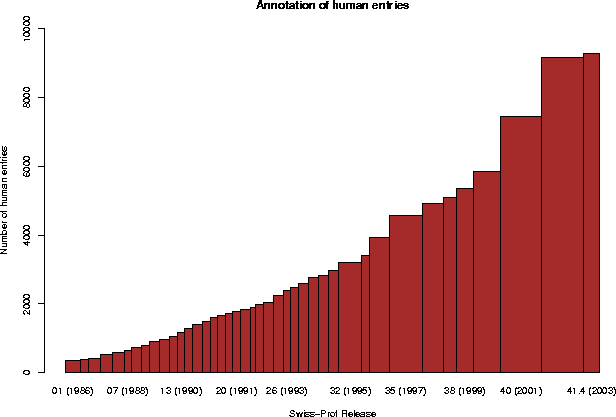 4. SEQUENCE SIZE
Repartition of the sequences by size (excluding fragments)
From To Number From To Number
1- 50 2287 1001-1100 1141
51- 100 8563 1101-1200 812
101- 150 12707 1201-1300 573
151- 200 11465 1301-1400 387
201- 250 11626 1401-1500 312
251- 300 10218 1501-1600 217
301- 350 10234 1601-1700 168
351- 400 9944 1701-1800 120
401- 450 7585 1801-1900 129
451- 500 6631 1901-2000 108
501- 550 5184 2001-2100 59
551- 600 3439 2101-2200 97
601- 650 2786 2201-2300 99
651- 700 2031 2301-2400 57
701- 750 1777 2401-2500 56
751- 800 1486 >2500 330
801- 850 1122
851- 900 1163
901- 950 830
951-1000 712
4. SEQUENCE SIZE
Repartition of the sequences by size (excluding fragments)
From To Number From To Number
1- 50 2287 1001-1100 1141
51- 100 8563 1101-1200 812
101- 150 12707 1201-1300 573
151- 200 11465 1301-1400 387
201- 250 11626 1401-1500 312
251- 300 10218 1501-1600 217
301- 350 10234 1601-1700 168
351- 400 9944 1701-1800 120
401- 450 7585 1801-1900 129
451- 500 6631 1901-2000 108
501- 550 5184 2001-2100 59
551- 600 3439 2101-2200 97
601- 650 2786 2201-2300 99
651- 700 2031 2301-2400 57
701- 750 1777 2401-2500 56
751- 800 1486 >2500 330
801- 850 1122
851- 900 1163
901- 950 830
951-1000 712
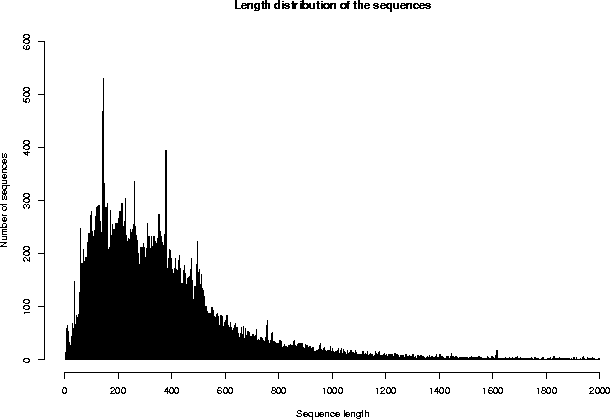 The average sequence length in Swiss-Prot is 367 amino acids.
The shortest sequence is GRWM_HUMAN (P01157): 3 amino acids.
The longest sequence is SNE1_HUMAN (Q8NF91): 8797 amino acids.
5. JOURNAL CITATIONS
Note: the following citation statistics reflect the number of distinct
journal citations.
Total number of journals cited in this release of Swiss-Prot: 1325
5.1 Table of the frequency of journal citations
Journals cited 1x: 498
2x: 167
3x: 89
4x: 60
5x: 49
6x: 41
7x: 26
8x: 27
9x: 21
10x: 10
11- 20x: 102
21- 50x: 99
51-100x: 40
>100x: 96
5.2 List of the most cited journals in Swiss-Prot
Nb Citations Journal name
-- --------- -------------------------------------------------------------
1 9201 Journal of Biological Chemistry
2 5039 Proceedings of the National Academy of Sciences of the U.S.A.
3 3641 Nucleic Acids Research
4 3631 Journal of Bacteriology
5 3401 Gene
6 2675 FEBS Letters
7 2614 Biochemical and Biophysical Research Communications
8 2436 European Journal of Biochemistry
9 2393 Biochemistry
10 2180 The EMBO Journal
11 2056 Nature
12 2033 Biochimica et Biophysica Acta
13 1829 Journal of Molecular Biology
14 1764 Genomics
15 1598 Cell
16 1560 Molecular and Cellular Biology
17 1257 Biochemical Journal
18 1167 Science
19 1124 Plant Molecular Biology
20 1118 Molecular and General Genetics
21 1069 Molecular Microbiology
22 858 Journal of Biochemistry
23 832 Virology
24 754 Human Molecular Genetics
25 702 Journal of Cell Biology
26 652 Nature Genetics
27 600 Journal of Virology
28 593 Plant Physiology
29 590 Genes and Development
30 582 Human Mutation
31 554 Oncogene
32 542 The American Journal of Human Genetics
33 531 Infection and Immunity
34 530 Yeast
35 520 Journal of Immunology
36 498 Journal of General Virology
37 474 Archives of Biochemistry and Biophysics
38 456 Structure
39 447 FEMS Microbiology Letters
40 433 Microbiology
41 408 Development
42 383 Human Genetics
43 380 Nature Structural Biology
44 377 Current Genetics
45 352 Genetics
46 350 Molecular and Biochemical Parasitology
47 336 Blood
48 318 Applied and Environmental Microbiology
49 314 Journal of Clinical Investigation
50 303 Molecular Endocrinology
51 284 DNA and Cell Biology
52 284 Mammalian Genome
53 283 Journal of Molecular Evolution
54 282 Protein Science
55 279 Developmental Biology
56 271 Biological Chemistry Hoppe-Seyler
57 256 Cancer Research
58 249 Journal of Experimental Medicine
59 248 Neuron
60 244 Immunogenetics
61 244 Mechanisms of Development
62 231 Endocrinology
63 229 Journal of General Microbiology
64 223 DNA Sequence
65 219 Acta Crystallographica, Section D
66 215 The Plant Cell
67 213 Hoppe-Seyler's Zeitschrift fur Physiologische Chemie
68 212 Molecular Biology of the Cell
69 210 Journal of Cell Science
70 193 Molecular Biology and Evolution
71 192 Brain Research. Molecular Brain Research
72 189 The Plant Journal
73 185 Journal of Neurochemistry
74 183 Journal of Neuroscience
75 161 Comparative Biochemistry and Physiology
76 160 Cytogenetics and Cell Genetics
77 156 DNA
78 155 The Journal of Clinical Endocrinology and Metabolism
79 155 Bioscience, Biotechnology, and Biochemistry
80 150 Molecular Pharmacology
81 145 Toxicon
82 144 Antimicrobial Agents and Chemotherapy
83 141 American Journal of Physiology
84 131 Biochimie
85 127 Bioorganicheskaia Khimiia
86 125 Proteins
87 125 Virus Research
88 124 DNA Research
89 123 Molecular Plant-Microbe Interactions
90 119 Hemoglobin
91 117 Peptides
92 115 Current Biology
93 114 Agricultural and Biological Chemistry
94 112 Journal of Investigative Dermatology
95 111 Molecular and Cellular Endocrinology
96 106 Genome Research
97 100 Molecular Cell
6. STATISTICS FOR SOME LINE TYPES
The following table summarizes the total number of some Swiss-Prot lines,
as well as the number of entries with at least one such line, and the
frequency of the lines.
Total Number of Average
Line type / subtype number entries per entry
--------------------------------- -------- --------- ---------
References (RL) 236510 1.90
Journal 202609 114005 1.63
Submitted to EMBL/GenBank/DDBJ 31371 26413 0.25
Unpublished observations 535 531 <0.01
Plant Gene Register 468 458 <0.01
Submitted to Swiss-Prot 468 466 <0.01
Book citation 460 450 <0.01
Thesis 192 190 <0.01
Submitted to other databases 190 189 <0.01
Unpublished results 123 121 <0.01
Patent 92 91 <0.01
Worm Breeder's Gazette 2 2 <0.01
Comments (CC) 414220 3.33
SIMILARITY 120602 105431 0.97
FUNCTION 78373 77063 0.63
SUBCELLULAR LOCATION 56267 56267 0.45
CATALYTIC ACTIVITY 40450 38045 0.32
SUBUNIT 34492 34492 0.28
PATHWAY 18129 17598 0.15
TISSUE SPECIFICITY 13929 13929 0.11
COFACTOR 12482 12482 0.10
MISCELLANEOUS 7944 7312 0.06
PTM 7243 6648 0.06
ALTERNATIVE PRODUCTS 4037 4037 0.03
DOMAIN 3613 3320 0.03
INDUCTION 3606 3606 0.03
CAUTION 3444 3261 0.03
DEVELOPMENTAL STAGE 3395 3395 0.03
DISEASE 2265 1877 0.02
ENZYME REGULATION 1766 1766 0.01
MASS SPECTROMETRY 896 813 0.01
DATABASE 842 775 0.01
POLYMORPHISM 348 339 <0.01
BIOTECHNOLOGY 50 50 <0.01
PHARMACEUTICAL 47 47 <0.01
Features (FT) 664824 5.34
DOMAIN 97486 29473 0.78
TRANSMEM 78072 17192 0.63
CONFLICT 48130 16912 0.39
CARBOHYD 45883 11232 0.37
DISULFID 42140 10973 0.34
TURN 39177 2956 0.31
METAL 38095 10271 0.31
STRAND 36304 2644 0.29
HELIX 27742 2845 0.22
ACT_SITE 24809 15515 0.20
VARIANT 23819 4458 0.19
CHAIN 23600 19302 0.19
REPEAT 22771 3776 0.18
NP_BIND 15790 11115 0.13
SIGNAL 14915 14913 0.12
MOD_RES 13433 7558 0.11
NON_TER 10331 7882 0.08
BINDING 8202 6348 0.07
ZN_FING 7875 2806 0.06
VARSPLIC 7174 3336 0.06
SITE 6444 4425 0.05
INIT_MET 5628 5591 0.05
PROPEP 4748 4059 0.04
MUTAGEN 4485 1386 0.04
DNA_BIND 4284 4030 0.03
CA_BIND 4051 1150 0.03
LIPID 2955 2400 0.02
TRANSIT 2617 2596 0.02
PEPTIDE 2534 1017 0.02
NON_CONS 806 413 0.01
UNSURE 290 123 <0.01
SE_CYS 116 78 <0.01
THIOETH 94 32 <0.01
THIOLEST 24 24 <0.01
Cross-references (DR) 1039217 8.35
EMBL 234705 118155 1.89
InterPro 197565 105320 1.59
Pfam 134362 100588 1.08
PROSITE 107203 67869 0.86
PIR 47040 35734 0.38
PRINTS 39696 35082 0.32
SMART 39012 29665 0.31
HSSP 38193 38193 0.31
GO 36337 12037 0.29
TIGRFAMs 31733 29373 0.25
ProDom 30371 29067 0.24
HAMAP 25218 25122 0.20
PDB 11817 3567 0.09
TIGR 11289 11243 0.09
MIM 8235 7136 0.07
Genew 7947 7899 0.06
MGD 5909 5894 0.05
SGD 4937 4883 0.04
EcoGene 4228 4226 0.03
MEROPS 3325 3231 0.03
TRANSFAC 2474 2224 0.02
WormPep 2466 2277 0.02
SubtiList 2366 2365 0.02
FlyBase 2275 2212 0.02
GeneDB_SPombe 2145 2115 0.02
TubercuList 1404 1367 0.01
StyGene 1213 1210 0.01
SWISS-2DPAGE 810 809 0.01
ListiList 728 673 0.01
Leproma 590 586 <0.01
Gramene 414 412 <0.01
MaizeDB 405 401 <0.01
HIV 370 354 <0.01
REBASE 358 353 <0.01
ECO2DBASE 351 299 <0.01
DictyDb 319 316 <0.01
GlycoSuiteDB 259 259 <0.01
ZFIN 229 229 <0.01
PHCI-2DPAGE 211 211 <0.01
MypuList 135 135 <0.01
Aarhus/Ghent-2DPAGE 128 98 <0.01
Siena-2DPAGE 104 104 <0.01
HSC-2DPAGE 85 85 <0.01
PhosSite 53 53 <0.01
SagaList 52 52 <0.01
COMPLUYEAST-2DPAGE 50 50 <0.01
PMMA-2DPAGE 47 47 <0.01
Maize-2DPAGE 39 39 <0.01
ANU-2DPAGE 15 15 <0.01
7. MISCELLANEOUS STATISTICS
Total number of distinct authors cited in Swiss-Prot: 165729
Total number of entries encoded on a chloroplast: 3185
Total number of entries encoded on a mitochondrion: 2386
Total number of entries encoded on a cyanelle: 145
Total number of entries encoded on a plasmid: 2643
Number of fragments: 8009
Number of additional sequences encoded on splice variants: 5845
The average sequence length in Swiss-Prot is 367 amino acids.
The shortest sequence is GRWM_HUMAN (P01157): 3 amino acids.
The longest sequence is SNE1_HUMAN (Q8NF91): 8797 amino acids.
5. JOURNAL CITATIONS
Note: the following citation statistics reflect the number of distinct
journal citations.
Total number of journals cited in this release of Swiss-Prot: 1325
5.1 Table of the frequency of journal citations
Journals cited 1x: 498
2x: 167
3x: 89
4x: 60
5x: 49
6x: 41
7x: 26
8x: 27
9x: 21
10x: 10
11- 20x: 102
21- 50x: 99
51-100x: 40
>100x: 96
5.2 List of the most cited journals in Swiss-Prot
Nb Citations Journal name
-- --------- -------------------------------------------------------------
1 9201 Journal of Biological Chemistry
2 5039 Proceedings of the National Academy of Sciences of the U.S.A.
3 3641 Nucleic Acids Research
4 3631 Journal of Bacteriology
5 3401 Gene
6 2675 FEBS Letters
7 2614 Biochemical and Biophysical Research Communications
8 2436 European Journal of Biochemistry
9 2393 Biochemistry
10 2180 The EMBO Journal
11 2056 Nature
12 2033 Biochimica et Biophysica Acta
13 1829 Journal of Molecular Biology
14 1764 Genomics
15 1598 Cell
16 1560 Molecular and Cellular Biology
17 1257 Biochemical Journal
18 1167 Science
19 1124 Plant Molecular Biology
20 1118 Molecular and General Genetics
21 1069 Molecular Microbiology
22 858 Journal of Biochemistry
23 832 Virology
24 754 Human Molecular Genetics
25 702 Journal of Cell Biology
26 652 Nature Genetics
27 600 Journal of Virology
28 593 Plant Physiology
29 590 Genes and Development
30 582 Human Mutation
31 554 Oncogene
32 542 The American Journal of Human Genetics
33 531 Infection and Immunity
34 530 Yeast
35 520 Journal of Immunology
36 498 Journal of General Virology
37 474 Archives of Biochemistry and Biophysics
38 456 Structure
39 447 FEMS Microbiology Letters
40 433 Microbiology
41 408 Development
42 383 Human Genetics
43 380 Nature Structural Biology
44 377 Current Genetics
45 352 Genetics
46 350 Molecular and Biochemical Parasitology
47 336 Blood
48 318 Applied and Environmental Microbiology
49 314 Journal of Clinical Investigation
50 303 Molecular Endocrinology
51 284 DNA and Cell Biology
52 284 Mammalian Genome
53 283 Journal of Molecular Evolution
54 282 Protein Science
55 279 Developmental Biology
56 271 Biological Chemistry Hoppe-Seyler
57 256 Cancer Research
58 249 Journal of Experimental Medicine
59 248 Neuron
60 244 Immunogenetics
61 244 Mechanisms of Development
62 231 Endocrinology
63 229 Journal of General Microbiology
64 223 DNA Sequence
65 219 Acta Crystallographica, Section D
66 215 The Plant Cell
67 213 Hoppe-Seyler's Zeitschrift fur Physiologische Chemie
68 212 Molecular Biology of the Cell
69 210 Journal of Cell Science
70 193 Molecular Biology and Evolution
71 192 Brain Research. Molecular Brain Research
72 189 The Plant Journal
73 185 Journal of Neurochemistry
74 183 Journal of Neuroscience
75 161 Comparative Biochemistry and Physiology
76 160 Cytogenetics and Cell Genetics
77 156 DNA
78 155 The Journal of Clinical Endocrinology and Metabolism
79 155 Bioscience, Biotechnology, and Biochemistry
80 150 Molecular Pharmacology
81 145 Toxicon
82 144 Antimicrobial Agents and Chemotherapy
83 141 American Journal of Physiology
84 131 Biochimie
85 127 Bioorganicheskaia Khimiia
86 125 Proteins
87 125 Virus Research
88 124 DNA Research
89 123 Molecular Plant-Microbe Interactions
90 119 Hemoglobin
91 117 Peptides
92 115 Current Biology
93 114 Agricultural and Biological Chemistry
94 112 Journal of Investigative Dermatology
95 111 Molecular and Cellular Endocrinology
96 106 Genome Research
97 100 Molecular Cell
6. STATISTICS FOR SOME LINE TYPES
The following table summarizes the total number of some Swiss-Prot lines,
as well as the number of entries with at least one such line, and the
frequency of the lines.
Total Number of Average
Line type / subtype number entries per entry
--------------------------------- -------- --------- ---------
References (RL) 236510 1.90
Journal 202609 114005 1.63
Submitted to EMBL/GenBank/DDBJ 31371 26413 0.25
Unpublished observations 535 531 <0.01
Plant Gene Register 468 458 <0.01
Submitted to Swiss-Prot 468 466 <0.01
Book citation 460 450 <0.01
Thesis 192 190 <0.01
Submitted to other databases 190 189 <0.01
Unpublished results 123 121 <0.01
Patent 92 91 <0.01
Worm Breeder's Gazette 2 2 <0.01
Comments (CC) 414220 3.33
SIMILARITY 120602 105431 0.97
FUNCTION 78373 77063 0.63
SUBCELLULAR LOCATION 56267 56267 0.45
CATALYTIC ACTIVITY 40450 38045 0.32
SUBUNIT 34492 34492 0.28
PATHWAY 18129 17598 0.15
TISSUE SPECIFICITY 13929 13929 0.11
COFACTOR 12482 12482 0.10
MISCELLANEOUS 7944 7312 0.06
PTM 7243 6648 0.06
ALTERNATIVE PRODUCTS 4037 4037 0.03
DOMAIN 3613 3320 0.03
INDUCTION 3606 3606 0.03
CAUTION 3444 3261 0.03
DEVELOPMENTAL STAGE 3395 3395 0.03
DISEASE 2265 1877 0.02
ENZYME REGULATION 1766 1766 0.01
MASS SPECTROMETRY 896 813 0.01
DATABASE 842 775 0.01
POLYMORPHISM 348 339 <0.01
BIOTECHNOLOGY 50 50 <0.01
PHARMACEUTICAL 47 47 <0.01
Features (FT) 664824 5.34
DOMAIN 97486 29473 0.78
TRANSMEM 78072 17192 0.63
CONFLICT 48130 16912 0.39
CARBOHYD 45883 11232 0.37
DISULFID 42140 10973 0.34
TURN 39177 2956 0.31
METAL 38095 10271 0.31
STRAND 36304 2644 0.29
HELIX 27742 2845 0.22
ACT_SITE 24809 15515 0.20
VARIANT 23819 4458 0.19
CHAIN 23600 19302 0.19
REPEAT 22771 3776 0.18
NP_BIND 15790 11115 0.13
SIGNAL 14915 14913 0.12
MOD_RES 13433 7558 0.11
NON_TER 10331 7882 0.08
BINDING 8202 6348 0.07
ZN_FING 7875 2806 0.06
VARSPLIC 7174 3336 0.06
SITE 6444 4425 0.05
INIT_MET 5628 5591 0.05
PROPEP 4748 4059 0.04
MUTAGEN 4485 1386 0.04
DNA_BIND 4284 4030 0.03
CA_BIND 4051 1150 0.03
LIPID 2955 2400 0.02
TRANSIT 2617 2596 0.02
PEPTIDE 2534 1017 0.02
NON_CONS 806 413 0.01
UNSURE 290 123 <0.01
SE_CYS 116 78 <0.01
THIOETH 94 32 <0.01
THIOLEST 24 24 <0.01
Cross-references (DR) 1039217 8.35
EMBL 234705 118155 1.89
InterPro 197565 105320 1.59
Pfam 134362 100588 1.08
PROSITE 107203 67869 0.86
PIR 47040 35734 0.38
PRINTS 39696 35082 0.32
SMART 39012 29665 0.31
HSSP 38193 38193 0.31
GO 36337 12037 0.29
TIGRFAMs 31733 29373 0.25
ProDom 30371 29067 0.24
HAMAP 25218 25122 0.20
PDB 11817 3567 0.09
TIGR 11289 11243 0.09
MIM 8235 7136 0.07
Genew 7947 7899 0.06
MGD 5909 5894 0.05
SGD 4937 4883 0.04
EcoGene 4228 4226 0.03
MEROPS 3325 3231 0.03
TRANSFAC 2474 2224 0.02
WormPep 2466 2277 0.02
SubtiList 2366 2365 0.02
FlyBase 2275 2212 0.02
GeneDB_SPombe 2145 2115 0.02
TubercuList 1404 1367 0.01
StyGene 1213 1210 0.01
SWISS-2DPAGE 810 809 0.01
ListiList 728 673 0.01
Leproma 590 586 <0.01
Gramene 414 412 <0.01
MaizeDB 405 401 <0.01
HIV 370 354 <0.01
REBASE 358 353 <0.01
ECO2DBASE 351 299 <0.01
DictyDb 319 316 <0.01
GlycoSuiteDB 259 259 <0.01
ZFIN 229 229 <0.01
PHCI-2DPAGE 211 211 <0.01
MypuList 135 135 <0.01
Aarhus/Ghent-2DPAGE 128 98 <0.01
Siena-2DPAGE 104 104 <0.01
HSC-2DPAGE 85 85 <0.01
PhosSite 53 53 <0.01
SagaList 52 52 <0.01
COMPLUYEAST-2DPAGE 50 50 <0.01
PMMA-2DPAGE 47 47 <0.01
Maize-2DPAGE 39 39 <0.01
ANU-2DPAGE 15 15 <0.01
7. MISCELLANEOUS STATISTICS
Total number of distinct authors cited in Swiss-Prot: 165729
Total number of entries encoded on a chloroplast: 3185
Total number of entries encoded on a mitochondrion: 2386
Total number of entries encoded on a cyanelle: 145
Total number of entries encoded on a plasmid: 2643
Number of fragments: 8009
Number of additional sequences encoded on splice variants: 5845
 ExPASy Home page
ExPASy Home page ExPASy Home page
ExPASy Home page 2. AMINO ACID COMPOSITION
2.1 Composition in percent for the complete database
Ala (A) 7.72 Gln (Q) 3.92 Leu (L) 9.57 Ser (S) 6.98
Arg (R) 5.24 Glu (E) 6.55 Lys (K) 5.97 Thr (T) 5.51
Asn (N) 4.28 Gly (G) 6.90 Met (M) 2.37 Trp (W) 1.18
Asp (D) 5.28 His (H) 2.26 Phe (F) 4.06 Tyr (Y) 3.12
Cys (C) 1.59 Ile (I) 5.89 Pro (P) 4.87 Val (V) 6.66
Asx (B) 0.000 Glx (Z) 0.000 Xaa (X) 0.01
2. AMINO ACID COMPOSITION
2.1 Composition in percent for the complete database
Ala (A) 7.72 Gln (Q) 3.92 Leu (L) 9.57 Ser (S) 6.98
Arg (R) 5.24 Glu (E) 6.55 Lys (K) 5.97 Thr (T) 5.51
Asn (N) 4.28 Gly (G) 6.90 Met (M) 2.37 Trp (W) 1.18
Asp (D) 5.28 His (H) 2.26 Phe (F) 4.06 Tyr (Y) 3.12
Cys (C) 1.59 Ile (I) 5.89 Pro (P) 4.87 Val (V) 6.66
Asx (B) 0.000 Glx (Z) 0.000 Xaa (X) 0.01
 Legend: gray = aliphatic, red = acidic, green = small hydroxy,
blue = basic, black = aromatic, white = amide, yellow = sulfur
2.2 Classification of the amino acids by their frequency
Leu, Ala, Ser, Gly, Val, Glu, Lys, Ile, Thr, Asp, Arg, Pro, Asn, Phe,
Gln, Tyr, Met, His, Cys, Trp
3. TAXONOMIC ORIGIN
Total number of species represented in this release of Swiss-Prot: 7830
The first twenty species represent 52260 sequences: 42 % of the total
number of entries.
3.1 Table of the frequency of occurrence of species
Species represented 1x: 3701
2x: 1210
3x: 631
4x: 406
5x: 261
6x: 256
7x: 198
8x: 147
9x: 124
10x: 66
11- 20x: 330
21- 50x: 248
51-100x: 91
>100x: 161
3.2 Table of the most represented species
------ --------- --------------------------------------------
Number Frequency Species
------ --------- --------------------------------------------
1 9294 Homo sapiens (Human)
2 6247 Mus musculus (Mouse)
3 4893 Saccharomyces cerevisiae (Baker's yeast)
4 4832 Escherichia coli
5 3465 Rattus norvegicus (Rat)
6 2406 Bacillus subtilis
7 2321 Caenorhabditis elegans
8 2130 Schizosaccharomyces pombe (Fission yeast)
9 2092 Arabidopsis thaliana (Mouse-ear cress)
10 1794 Drosophila melanogaster (Fruit fly)
11 1773 Haemophilus influenzae
12 1530 Methanococcus jannaschii
13 1506 Escherichia coli O157:H7
14 1394 Bos taurus (Bovine)
15 1375 Mycobacterium tuberculosis
16 1257 Salmonella typhimurium
17 1064 Gallus gallus (Chicken)
18 1006 Escherichia coli O6
19 961 Shigella flexneri
20 920 Synechocystis sp. (strain PCC 6803)
21 880 Archaeoglobus fulgidus
22 846 Pseudomonas aeruginosa
23 845 Xenopus laevis (African clawed frog)
24 823 Sus scrofa (Pig)
25 796 Salmonella typhi
26 716 Aquifex aeolicus
27 705 Oryctolagus cuniculus (Rabbit)
28 687 Mycoplasma pneumoniae
29 681 Rhizobium meliloti (Sinorhizobium meliloti)
30 624 Vibrio cholerae
31 599 Treponema pallidum
32 586 Mycobacterium leprae
33 572 Buchnera aphidicola (subsp. Acyrthosiphon pisum)
34 560 Buchnera aphidicola (subsp. Schizaphis graminum)
35 544 Yersinia pestis
36 537 Helicobacter pylori (Campylobacter pylori)
37 535 Rickettsia prowazekii
38 526 Streptomyces coelicolor
39 520 Helicobacter pylori J99 (Campylobacter pylori J99)
40 498 Bacillus halodurans
41 495 Methanobacterium thermoautotrophicum
42 491 Zea mays (Maize)
43 489 Pasteurella multocida
44 486 Mycoplasma genitalium
45 469 Anabaena sp. (strain PCC 7120)
46 437 Lactococcus lactis (subsp. lactis) (Streptococcus lactis)
47 421 Thermotoga maritima
48 417 Oryza sativa (Rice)
49 406 Borrelia burgdorferi (Lyme disease spirochete)
50 405 Chlamydia trachomatis
51 403 Rhizobium sp. (strain NGR234)
52 394 Canis familiaris (Dog)
53 394 Neisseria meningitidis (serogroup B)
54 392 Chlamydia pneumoniae (Chlamydophila pneumoniae)
55 390 Neisseria meningitidis (serogroup A)
56 382 Chlamydia muridarum
57 370 Pyrococcus horikoshii
58 369 Caulobacter crescentus
59 367 Listeria monocytogenes
60 367 Clostridium acetobutylicum
61 363 Pyrococcus abyssi
62 363 Rhizobium loti (Mesorhizobium loti)
63 362 Ralstonia solanacearum (Pseudomonas solanacearum)
64 360 Listeria innocua
65 358 Streptococcus pneumoniae
66 356 Agrobacterium tumefaciens (strain C58 / ATCC 33970)
67 343 Nicotiana tabacum (Common tobacco)
68 342 Xylella fastidiosa
69 339 Deinococcus radiodurans
70 335 Xanthomonas campestris (pv. campestris)
71 333 Ovis aries (Sheep)
72 327 Staphylococcus aureus (strain N315)
73 326 Halobacterium sp. (strain NRC-1 / ATCC 700922 / JCM 11081)
74 326 Campylobacter jejuni
75 324 Staphylococcus aureus (strain Mu50 / ATCC 700699)
76 319 Clostridium perfringens
77 316 Dictyostelium discoideum (Slime mold)
78 312 Corynebacterium glutamicum (Brevibacterium flavum)
79 306 Sulfolobus solfataricus
80 304 Staphylococcus aureus (strain MW2)
81 299 Xanthomonas axonopodis (pv. citri)
82 291 Streptococcus pyogenes
83 289 Aeropyrum pernix
84 289 Pisum sativum (Garden pea)
85 285 Pyrococcus furiosus
86 279 Staphylococcus aureus
87 276 Brucella melitensis
88 268 Bacteriophage T4
89 266 Neurospora crassa
90 265 Triticum aestivum (Wheat)
91 264 Rickettsia conorii
92 264 Candida albicans (Yeast)
93 263 Thermoanaerobacter tengcongensis
94 258 Hordeum vulgare (Barley)
95 254 Vaccinia virus (strain Copenhagen)
96 254 Methanosarcina mazei (Methanosarcina frisia)
97 254 Glycine max (Soybean)
98 253 Methanosarcina acetivorans
99 251 Lycopersicon esculentum (Tomato)
100 248 Rhodobacter capsulatus (Rhodopseudomonas capsulata)
3.3 Taxonomic distribution of the sequences
Legend: gray = aliphatic, red = acidic, green = small hydroxy,
blue = basic, black = aromatic, white = amide, yellow = sulfur
2.2 Classification of the amino acids by their frequency
Leu, Ala, Ser, Gly, Val, Glu, Lys, Ile, Thr, Asp, Arg, Pro, Asn, Phe,
Gln, Tyr, Met, His, Cys, Trp
3. TAXONOMIC ORIGIN
Total number of species represented in this release of Swiss-Prot: 7830
The first twenty species represent 52260 sequences: 42 % of the total
number of entries.
3.1 Table of the frequency of occurrence of species
Species represented 1x: 3701
2x: 1210
3x: 631
4x: 406
5x: 261
6x: 256
7x: 198
8x: 147
9x: 124
10x: 66
11- 20x: 330
21- 50x: 248
51-100x: 91
>100x: 161
3.2 Table of the most represented species
------ --------- --------------------------------------------
Number Frequency Species
------ --------- --------------------------------------------
1 9294 Homo sapiens (Human)
2 6247 Mus musculus (Mouse)
3 4893 Saccharomyces cerevisiae (Baker's yeast)
4 4832 Escherichia coli
5 3465 Rattus norvegicus (Rat)
6 2406 Bacillus subtilis
7 2321 Caenorhabditis elegans
8 2130 Schizosaccharomyces pombe (Fission yeast)
9 2092 Arabidopsis thaliana (Mouse-ear cress)
10 1794 Drosophila melanogaster (Fruit fly)
11 1773 Haemophilus influenzae
12 1530 Methanococcus jannaschii
13 1506 Escherichia coli O157:H7
14 1394 Bos taurus (Bovine)
15 1375 Mycobacterium tuberculosis
16 1257 Salmonella typhimurium
17 1064 Gallus gallus (Chicken)
18 1006 Escherichia coli O6
19 961 Shigella flexneri
20 920 Synechocystis sp. (strain PCC 6803)
21 880 Archaeoglobus fulgidus
22 846 Pseudomonas aeruginosa
23 845 Xenopus laevis (African clawed frog)
24 823 Sus scrofa (Pig)
25 796 Salmonella typhi
26 716 Aquifex aeolicus
27 705 Oryctolagus cuniculus (Rabbit)
28 687 Mycoplasma pneumoniae
29 681 Rhizobium meliloti (Sinorhizobium meliloti)
30 624 Vibrio cholerae
31 599 Treponema pallidum
32 586 Mycobacterium leprae
33 572 Buchnera aphidicola (subsp. Acyrthosiphon pisum)
34 560 Buchnera aphidicola (subsp. Schizaphis graminum)
35 544 Yersinia pestis
36 537 Helicobacter pylori (Campylobacter pylori)
37 535 Rickettsia prowazekii
38 526 Streptomyces coelicolor
39 520 Helicobacter pylori J99 (Campylobacter pylori J99)
40 498 Bacillus halodurans
41 495 Methanobacterium thermoautotrophicum
42 491 Zea mays (Maize)
43 489 Pasteurella multocida
44 486 Mycoplasma genitalium
45 469 Anabaena sp. (strain PCC 7120)
46 437 Lactococcus lactis (subsp. lactis) (Streptococcus lactis)
47 421 Thermotoga maritima
48 417 Oryza sativa (Rice)
49 406 Borrelia burgdorferi (Lyme disease spirochete)
50 405 Chlamydia trachomatis
51 403 Rhizobium sp. (strain NGR234)
52 394 Canis familiaris (Dog)
53 394 Neisseria meningitidis (serogroup B)
54 392 Chlamydia pneumoniae (Chlamydophila pneumoniae)
55 390 Neisseria meningitidis (serogroup A)
56 382 Chlamydia muridarum
57 370 Pyrococcus horikoshii
58 369 Caulobacter crescentus
59 367 Listeria monocytogenes
60 367 Clostridium acetobutylicum
61 363 Pyrococcus abyssi
62 363 Rhizobium loti (Mesorhizobium loti)
63 362 Ralstonia solanacearum (Pseudomonas solanacearum)
64 360 Listeria innocua
65 358 Streptococcus pneumoniae
66 356 Agrobacterium tumefaciens (strain C58 / ATCC 33970)
67 343 Nicotiana tabacum (Common tobacco)
68 342 Xylella fastidiosa
69 339 Deinococcus radiodurans
70 335 Xanthomonas campestris (pv. campestris)
71 333 Ovis aries (Sheep)
72 327 Staphylococcus aureus (strain N315)
73 326 Halobacterium sp. (strain NRC-1 / ATCC 700922 / JCM 11081)
74 326 Campylobacter jejuni
75 324 Staphylococcus aureus (strain Mu50 / ATCC 700699)
76 319 Clostridium perfringens
77 316 Dictyostelium discoideum (Slime mold)
78 312 Corynebacterium glutamicum (Brevibacterium flavum)
79 306 Sulfolobus solfataricus
80 304 Staphylococcus aureus (strain MW2)
81 299 Xanthomonas axonopodis (pv. citri)
82 291 Streptococcus pyogenes
83 289 Aeropyrum pernix
84 289 Pisum sativum (Garden pea)
85 285 Pyrococcus furiosus
86 279 Staphylococcus aureus
87 276 Brucella melitensis
88 268 Bacteriophage T4
89 266 Neurospora crassa
90 265 Triticum aestivum (Wheat)
91 264 Rickettsia conorii
92 264 Candida albicans (Yeast)
93 263 Thermoanaerobacter tengcongensis
94 258 Hordeum vulgare (Barley)
95 254 Vaccinia virus (strain Copenhagen)
96 254 Methanosarcina mazei (Methanosarcina frisia)
97 254 Glycine max (Soybean)
98 253 Methanosarcina acetivorans
99 251 Lycopersicon esculentum (Tomato)
100 248 Rhodobacter capsulatus (Rhodopseudomonas capsulata)
3.3 Taxonomic distribution of the sequences
 Kingdom sequences (% of the database)
Archaea 7218 ( 6%)
Bacteria 47533 ( 38%)
Eukaryota 61240 ( 49%)
Viruses 8473 ( 7%)
Within Eukaryota:
Kingdom sequences (% of the database)
Archaea 7218 ( 6%)
Bacteria 47533 ( 38%)
Eukaryota 61240 ( 49%)
Viruses 8473 ( 7%)
Within Eukaryota:
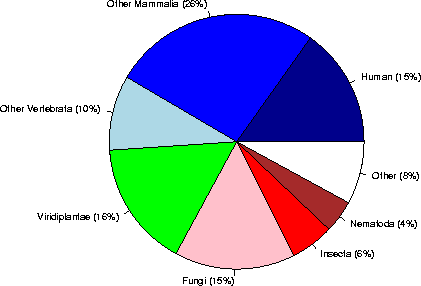 Category sequences (% of Eukaryota) (% of the complete database)
Human 9294 ( 15%) ( 7%)
Other Mammalia 16152 ( 26%) ( 13%)
Other Vertebrata 5824 ( 10%) ( 5%)
Viridiplantae 9814 ( 16%) ( 8%)
Fungi 9364 ( 15%) ( 8%)
Insecta 3394 ( 6%) ( 3%)
Nematoda 2535 ( 4%) ( 2%)
Other 4863 ( 8%) ( 4%)
3.4 Annotation of high-priority organisms
Category sequences (% of Eukaryota) (% of the complete database)
Human 9294 ( 15%) ( 7%)
Other Mammalia 16152 ( 26%) ( 13%)
Other Vertebrata 5824 ( 10%) ( 5%)
Viridiplantae 9814 ( 16%) ( 8%)
Fungi 9364 ( 15%) ( 8%)
Insecta 3394 ( 6%) ( 3%)
Nematoda 2535 ( 4%) ( 2%)
Other 4863 ( 8%) ( 4%)
3.4 Annotation of high-priority organisms

 4. SEQUENCE SIZE
Repartition of the sequences by size (excluding fragments)
From To Number From To Number
1- 50 2287 1001-1100 1141
51- 100 8563 1101-1200 812
101- 150 12707 1201-1300 573
151- 200 11465 1301-1400 387
201- 250 11626 1401-1500 312
251- 300 10218 1501-1600 217
301- 350 10234 1601-1700 168
351- 400 9944 1701-1800 120
401- 450 7585 1801-1900 129
451- 500 6631 1901-2000 108
501- 550 5184 2001-2100 59
551- 600 3439 2101-2200 97
601- 650 2786 2201-2300 99
651- 700 2031 2301-2400 57
701- 750 1777 2401-2500 56
751- 800 1486 >2500 330
801- 850 1122
851- 900 1163
901- 950 830
951-1000 712
4. SEQUENCE SIZE
Repartition of the sequences by size (excluding fragments)
From To Number From To Number
1- 50 2287 1001-1100 1141
51- 100 8563 1101-1200 812
101- 150 12707 1201-1300 573
151- 200 11465 1301-1400 387
201- 250 11626 1401-1500 312
251- 300 10218 1501-1600 217
301- 350 10234 1601-1700 168
351- 400 9944 1701-1800 120
401- 450 7585 1801-1900 129
451- 500 6631 1901-2000 108
501- 550 5184 2001-2100 59
551- 600 3439 2101-2200 97
601- 650 2786 2201-2300 99
651- 700 2031 2301-2400 57
701- 750 1777 2401-2500 56
751- 800 1486 >2500 330
801- 850 1122
851- 900 1163
901- 950 830
951-1000 712
 The average sequence length in Swiss-Prot is 367 amino acids.
The shortest sequence is GRWM_HUMAN (P01157): 3 amino acids.
The longest sequence is SNE1_HUMAN (Q8NF91): 8797 amino acids.
5. JOURNAL CITATIONS
Note: the following citation statistics reflect the number of distinct
journal citations.
Total number of journals cited in this release of Swiss-Prot: 1325
5.1 Table of the frequency of journal citations
Journals cited 1x: 498
2x: 167
3x: 89
4x: 60
5x: 49
6x: 41
7x: 26
8x: 27
9x: 21
10x: 10
11- 20x: 102
21- 50x: 99
51-100x: 40
>100x: 96
5.2 List of the most cited journals in Swiss-Prot
Nb Citations Journal name
-- --------- -------------------------------------------------------------
1 9201 Journal of Biological Chemistry
2 5039 Proceedings of the National Academy of Sciences of the U.S.A.
3 3641 Nucleic Acids Research
4 3631 Journal of Bacteriology
5 3401 Gene
6 2675 FEBS Letters
7 2614 Biochemical and Biophysical Research Communications
8 2436 European Journal of Biochemistry
9 2393 Biochemistry
10 2180 The EMBO Journal
11 2056 Nature
12 2033 Biochimica et Biophysica Acta
13 1829 Journal of Molecular Biology
14 1764 Genomics
15 1598 Cell
16 1560 Molecular and Cellular Biology
17 1257 Biochemical Journal
18 1167 Science
19 1124 Plant Molecular Biology
20 1118 Molecular and General Genetics
21 1069 Molecular Microbiology
22 858 Journal of Biochemistry
23 832 Virology
24 754 Human Molecular Genetics
25 702 Journal of Cell Biology
26 652 Nature Genetics
27 600 Journal of Virology
28 593 Plant Physiology
29 590 Genes and Development
30 582 Human Mutation
31 554 Oncogene
32 542 The American Journal of Human Genetics
33 531 Infection and Immunity
34 530 Yeast
35 520 Journal of Immunology
36 498 Journal of General Virology
37 474 Archives of Biochemistry and Biophysics
38 456 Structure
39 447 FEMS Microbiology Letters
40 433 Microbiology
41 408 Development
42 383 Human Genetics
43 380 Nature Structural Biology
44 377 Current Genetics
45 352 Genetics
46 350 Molecular and Biochemical Parasitology
47 336 Blood
48 318 Applied and Environmental Microbiology
49 314 Journal of Clinical Investigation
50 303 Molecular Endocrinology
51 284 DNA and Cell Biology
52 284 Mammalian Genome
53 283 Journal of Molecular Evolution
54 282 Protein Science
55 279 Developmental Biology
56 271 Biological Chemistry Hoppe-Seyler
57 256 Cancer Research
58 249 Journal of Experimental Medicine
59 248 Neuron
60 244 Immunogenetics
61 244 Mechanisms of Development
62 231 Endocrinology
63 229 Journal of General Microbiology
64 223 DNA Sequence
65 219 Acta Crystallographica, Section D
66 215 The Plant Cell
67 213 Hoppe-Seyler's Zeitschrift fur Physiologische Chemie
68 212 Molecular Biology of the Cell
69 210 Journal of Cell Science
70 193 Molecular Biology and Evolution
71 192 Brain Research. Molecular Brain Research
72 189 The Plant Journal
73 185 Journal of Neurochemistry
74 183 Journal of Neuroscience
75 161 Comparative Biochemistry and Physiology
76 160 Cytogenetics and Cell Genetics
77 156 DNA
78 155 The Journal of Clinical Endocrinology and Metabolism
79 155 Bioscience, Biotechnology, and Biochemistry
80 150 Molecular Pharmacology
81 145 Toxicon
82 144 Antimicrobial Agents and Chemotherapy
83 141 American Journal of Physiology
84 131 Biochimie
85 127 Bioorganicheskaia Khimiia
86 125 Proteins
87 125 Virus Research
88 124 DNA Research
89 123 Molecular Plant-Microbe Interactions
90 119 Hemoglobin
91 117 Peptides
92 115 Current Biology
93 114 Agricultural and Biological Chemistry
94 112 Journal of Investigative Dermatology
95 111 Molecular and Cellular Endocrinology
96 106 Genome Research
97 100 Molecular Cell
6. STATISTICS FOR SOME LINE TYPES
The following table summarizes the total number of some Swiss-Prot lines,
as well as the number of entries with at least one such line, and the
frequency of the lines.
Total Number of Average
Line type / subtype number entries per entry
--------------------------------- -------- --------- ---------
References (RL) 236510 1.90
Journal 202609 114005 1.63
Submitted to EMBL/GenBank/DDBJ 31371 26413 0.25
Unpublished observations 535 531 <0.01
Plant Gene Register 468 458 <0.01
Submitted to Swiss-Prot 468 466 <0.01
Book citation 460 450 <0.01
Thesis 192 190 <0.01
Submitted to other databases 190 189 <0.01
Unpublished results 123 121 <0.01
Patent 92 91 <0.01
Worm Breeder's Gazette 2 2 <0.01
Comments (CC) 414220 3.33
SIMILARITY 120602 105431 0.97
FUNCTION 78373 77063 0.63
SUBCELLULAR LOCATION 56267 56267 0.45
CATALYTIC ACTIVITY 40450 38045 0.32
SUBUNIT 34492 34492 0.28
PATHWAY 18129 17598 0.15
TISSUE SPECIFICITY 13929 13929 0.11
COFACTOR 12482 12482 0.10
MISCELLANEOUS 7944 7312 0.06
PTM 7243 6648 0.06
ALTERNATIVE PRODUCTS 4037 4037 0.03
DOMAIN 3613 3320 0.03
INDUCTION 3606 3606 0.03
CAUTION 3444 3261 0.03
DEVELOPMENTAL STAGE 3395 3395 0.03
DISEASE 2265 1877 0.02
ENZYME REGULATION 1766 1766 0.01
MASS SPECTROMETRY 896 813 0.01
DATABASE 842 775 0.01
POLYMORPHISM 348 339 <0.01
BIOTECHNOLOGY 50 50 <0.01
PHARMACEUTICAL 47 47 <0.01
Features (FT) 664824 5.34
DOMAIN 97486 29473 0.78
TRANSMEM 78072 17192 0.63
CONFLICT 48130 16912 0.39
CARBOHYD 45883 11232 0.37
DISULFID 42140 10973 0.34
TURN 39177 2956 0.31
METAL 38095 10271 0.31
STRAND 36304 2644 0.29
HELIX 27742 2845 0.22
ACT_SITE 24809 15515 0.20
VARIANT 23819 4458 0.19
CHAIN 23600 19302 0.19
REPEAT 22771 3776 0.18
NP_BIND 15790 11115 0.13
SIGNAL 14915 14913 0.12
MOD_RES 13433 7558 0.11
NON_TER 10331 7882 0.08
BINDING 8202 6348 0.07
ZN_FING 7875 2806 0.06
VARSPLIC 7174 3336 0.06
SITE 6444 4425 0.05
INIT_MET 5628 5591 0.05
PROPEP 4748 4059 0.04
MUTAGEN 4485 1386 0.04
DNA_BIND 4284 4030 0.03
CA_BIND 4051 1150 0.03
LIPID 2955 2400 0.02
TRANSIT 2617 2596 0.02
PEPTIDE 2534 1017 0.02
NON_CONS 806 413 0.01
UNSURE 290 123 <0.01
SE_CYS 116 78 <0.01
THIOETH 94 32 <0.01
THIOLEST 24 24 <0.01
Cross-references (DR) 1039217 8.35
EMBL 234705 118155 1.89
InterPro 197565 105320 1.59
Pfam 134362 100588 1.08
PROSITE 107203 67869 0.86
PIR 47040 35734 0.38
PRINTS 39696 35082 0.32
SMART 39012 29665 0.31
HSSP 38193 38193 0.31
GO 36337 12037 0.29
TIGRFAMs 31733 29373 0.25
ProDom 30371 29067 0.24
HAMAP 25218 25122 0.20
PDB 11817 3567 0.09
TIGR 11289 11243 0.09
MIM 8235 7136 0.07
Genew 7947 7899 0.06
MGD 5909 5894 0.05
SGD 4937 4883 0.04
EcoGene 4228 4226 0.03
MEROPS 3325 3231 0.03
TRANSFAC 2474 2224 0.02
WormPep 2466 2277 0.02
SubtiList 2366 2365 0.02
FlyBase 2275 2212 0.02
GeneDB_SPombe 2145 2115 0.02
TubercuList 1404 1367 0.01
StyGene 1213 1210 0.01
SWISS-2DPAGE 810 809 0.01
ListiList 728 673 0.01
Leproma 590 586 <0.01
Gramene 414 412 <0.01
MaizeDB 405 401 <0.01
HIV 370 354 <0.01
REBASE 358 353 <0.01
ECO2DBASE 351 299 <0.01
DictyDb 319 316 <0.01
GlycoSuiteDB 259 259 <0.01
ZFIN 229 229 <0.01
PHCI-2DPAGE 211 211 <0.01
MypuList 135 135 <0.01
Aarhus/Ghent-2DPAGE 128 98 <0.01
Siena-2DPAGE 104 104 <0.01
HSC-2DPAGE 85 85 <0.01
PhosSite 53 53 <0.01
SagaList 52 52 <0.01
COMPLUYEAST-2DPAGE 50 50 <0.01
PMMA-2DPAGE 47 47 <0.01
Maize-2DPAGE 39 39 <0.01
ANU-2DPAGE 15 15 <0.01
7. MISCELLANEOUS STATISTICS
Total number of distinct authors cited in Swiss-Prot: 165729
Total number of entries encoded on a chloroplast: 3185
Total number of entries encoded on a mitochondrion: 2386
Total number of entries encoded on a cyanelle: 145
Total number of entries encoded on a plasmid: 2643
Number of fragments: 8009
Number of additional sequences encoded on splice variants: 5845
The average sequence length in Swiss-Prot is 367 amino acids.
The shortest sequence is GRWM_HUMAN (P01157): 3 amino acids.
The longest sequence is SNE1_HUMAN (Q8NF91): 8797 amino acids.
5. JOURNAL CITATIONS
Note: the following citation statistics reflect the number of distinct
journal citations.
Total number of journals cited in this release of Swiss-Prot: 1325
5.1 Table of the frequency of journal citations
Journals cited 1x: 498
2x: 167
3x: 89
4x: 60
5x: 49
6x: 41
7x: 26
8x: 27
9x: 21
10x: 10
11- 20x: 102
21- 50x: 99
51-100x: 40
>100x: 96
5.2 List of the most cited journals in Swiss-Prot
Nb Citations Journal name
-- --------- -------------------------------------------------------------
1 9201 Journal of Biological Chemistry
2 5039 Proceedings of the National Academy of Sciences of the U.S.A.
3 3641 Nucleic Acids Research
4 3631 Journal of Bacteriology
5 3401 Gene
6 2675 FEBS Letters
7 2614 Biochemical and Biophysical Research Communications
8 2436 European Journal of Biochemistry
9 2393 Biochemistry
10 2180 The EMBO Journal
11 2056 Nature
12 2033 Biochimica et Biophysica Acta
13 1829 Journal of Molecular Biology
14 1764 Genomics
15 1598 Cell
16 1560 Molecular and Cellular Biology
17 1257 Biochemical Journal
18 1167 Science
19 1124 Plant Molecular Biology
20 1118 Molecular and General Genetics
21 1069 Molecular Microbiology
22 858 Journal of Biochemistry
23 832 Virology
24 754 Human Molecular Genetics
25 702 Journal of Cell Biology
26 652 Nature Genetics
27 600 Journal of Virology
28 593 Plant Physiology
29 590 Genes and Development
30 582 Human Mutation
31 554 Oncogene
32 542 The American Journal of Human Genetics
33 531 Infection and Immunity
34 530 Yeast
35 520 Journal of Immunology
36 498 Journal of General Virology
37 474 Archives of Biochemistry and Biophysics
38 456 Structure
39 447 FEMS Microbiology Letters
40 433 Microbiology
41 408 Development
42 383 Human Genetics
43 380 Nature Structural Biology
44 377 Current Genetics
45 352 Genetics
46 350 Molecular and Biochemical Parasitology
47 336 Blood
48 318 Applied and Environmental Microbiology
49 314 Journal of Clinical Investigation
50 303 Molecular Endocrinology
51 284 DNA and Cell Biology
52 284 Mammalian Genome
53 283 Journal of Molecular Evolution
54 282 Protein Science
55 279 Developmental Biology
56 271 Biological Chemistry Hoppe-Seyler
57 256 Cancer Research
58 249 Journal of Experimental Medicine
59 248 Neuron
60 244 Immunogenetics
61 244 Mechanisms of Development
62 231 Endocrinology
63 229 Journal of General Microbiology
64 223 DNA Sequence
65 219 Acta Crystallographica, Section D
66 215 The Plant Cell
67 213 Hoppe-Seyler's Zeitschrift fur Physiologische Chemie
68 212 Molecular Biology of the Cell
69 210 Journal of Cell Science
70 193 Molecular Biology and Evolution
71 192 Brain Research. Molecular Brain Research
72 189 The Plant Journal
73 185 Journal of Neurochemistry
74 183 Journal of Neuroscience
75 161 Comparative Biochemistry and Physiology
76 160 Cytogenetics and Cell Genetics
77 156 DNA
78 155 The Journal of Clinical Endocrinology and Metabolism
79 155 Bioscience, Biotechnology, and Biochemistry
80 150 Molecular Pharmacology
81 145 Toxicon
82 144 Antimicrobial Agents and Chemotherapy
83 141 American Journal of Physiology
84 131 Biochimie
85 127 Bioorganicheskaia Khimiia
86 125 Proteins
87 125 Virus Research
88 124 DNA Research
89 123 Molecular Plant-Microbe Interactions
90 119 Hemoglobin
91 117 Peptides
92 115 Current Biology
93 114 Agricultural and Biological Chemistry
94 112 Journal of Investigative Dermatology
95 111 Molecular and Cellular Endocrinology
96 106 Genome Research
97 100 Molecular Cell
6. STATISTICS FOR SOME LINE TYPES
The following table summarizes the total number of some Swiss-Prot lines,
as well as the number of entries with at least one such line, and the
frequency of the lines.
Total Number of Average
Line type / subtype number entries per entry
--------------------------------- -------- --------- ---------
References (RL) 236510 1.90
Journal 202609 114005 1.63
Submitted to EMBL/GenBank/DDBJ 31371 26413 0.25
Unpublished observations 535 531 <0.01
Plant Gene Register 468 458 <0.01
Submitted to Swiss-Prot 468 466 <0.01
Book citation 460 450 <0.01
Thesis 192 190 <0.01
Submitted to other databases 190 189 <0.01
Unpublished results 123 121 <0.01
Patent 92 91 <0.01
Worm Breeder's Gazette 2 2 <0.01
Comments (CC) 414220 3.33
SIMILARITY 120602 105431 0.97
FUNCTION 78373 77063 0.63
SUBCELLULAR LOCATION 56267 56267 0.45
CATALYTIC ACTIVITY 40450 38045 0.32
SUBUNIT 34492 34492 0.28
PATHWAY 18129 17598 0.15
TISSUE SPECIFICITY 13929 13929 0.11
COFACTOR 12482 12482 0.10
MISCELLANEOUS 7944 7312 0.06
PTM 7243 6648 0.06
ALTERNATIVE PRODUCTS 4037 4037 0.03
DOMAIN 3613 3320 0.03
INDUCTION 3606 3606 0.03
CAUTION 3444 3261 0.03
DEVELOPMENTAL STAGE 3395 3395 0.03
DISEASE 2265 1877 0.02
ENZYME REGULATION 1766 1766 0.01
MASS SPECTROMETRY 896 813 0.01
DATABASE 842 775 0.01
POLYMORPHISM 348 339 <0.01
BIOTECHNOLOGY 50 50 <0.01
PHARMACEUTICAL 47 47 <0.01
Features (FT) 664824 5.34
DOMAIN 97486 29473 0.78
TRANSMEM 78072 17192 0.63
CONFLICT 48130 16912 0.39
CARBOHYD 45883 11232 0.37
DISULFID 42140 10973 0.34
TURN 39177 2956 0.31
METAL 38095 10271 0.31
STRAND 36304 2644 0.29
HELIX 27742 2845 0.22
ACT_SITE 24809 15515 0.20
VARIANT 23819 4458 0.19
CHAIN 23600 19302 0.19
REPEAT 22771 3776 0.18
NP_BIND 15790 11115 0.13
SIGNAL 14915 14913 0.12
MOD_RES 13433 7558 0.11
NON_TER 10331 7882 0.08
BINDING 8202 6348 0.07
ZN_FING 7875 2806 0.06
VARSPLIC 7174 3336 0.06
SITE 6444 4425 0.05
INIT_MET 5628 5591 0.05
PROPEP 4748 4059 0.04
MUTAGEN 4485 1386 0.04
DNA_BIND 4284 4030 0.03
CA_BIND 4051 1150 0.03
LIPID 2955 2400 0.02
TRANSIT 2617 2596 0.02
PEPTIDE 2534 1017 0.02
NON_CONS 806 413 0.01
UNSURE 290 123 <0.01
SE_CYS 116 78 <0.01
THIOETH 94 32 <0.01
THIOLEST 24 24 <0.01
Cross-references (DR) 1039217 8.35
EMBL 234705 118155 1.89
InterPro 197565 105320 1.59
Pfam 134362 100588 1.08
PROSITE 107203 67869 0.86
PIR 47040 35734 0.38
PRINTS 39696 35082 0.32
SMART 39012 29665 0.31
HSSP 38193 38193 0.31
GO 36337 12037 0.29
TIGRFAMs 31733 29373 0.25
ProDom 30371 29067 0.24
HAMAP 25218 25122 0.20
PDB 11817 3567 0.09
TIGR 11289 11243 0.09
MIM 8235 7136 0.07
Genew 7947 7899 0.06
MGD 5909 5894 0.05
SGD 4937 4883 0.04
EcoGene 4228 4226 0.03
MEROPS 3325 3231 0.03
TRANSFAC 2474 2224 0.02
WormPep 2466 2277 0.02
SubtiList 2366 2365 0.02
FlyBase 2275 2212 0.02
GeneDB_SPombe 2145 2115 0.02
TubercuList 1404 1367 0.01
StyGene 1213 1210 0.01
SWISS-2DPAGE 810 809 0.01
ListiList 728 673 0.01
Leproma 590 586 <0.01
Gramene 414 412 <0.01
MaizeDB 405 401 <0.01
HIV 370 354 <0.01
REBASE 358 353 <0.01
ECO2DBASE 351 299 <0.01
DictyDb 319 316 <0.01
GlycoSuiteDB 259 259 <0.01
ZFIN 229 229 <0.01
PHCI-2DPAGE 211 211 <0.01
MypuList 135 135 <0.01
Aarhus/Ghent-2DPAGE 128 98 <0.01
Siena-2DPAGE 104 104 <0.01
HSC-2DPAGE 85 85 <0.01
PhosSite 53 53 <0.01
SagaList 52 52 <0.01
COMPLUYEAST-2DPAGE 50 50 <0.01
PMMA-2DPAGE 47 47 <0.01
Maize-2DPAGE 39 39 <0.01
ANU-2DPAGE 15 15 <0.01
7. MISCELLANEOUS STATISTICS
Total number of distinct authors cited in Swiss-Prot: 165729
Total number of entries encoded on a chloroplast: 3185
Total number of entries encoded on a mitochondrion: 2386
Total number of entries encoded on a cyanelle: 145
Total number of entries encoded on a plasmid: 2643
Number of fragments: 8009
Number of additional sequences encoded on splice variants: 5845
 ExPASy Home page
ExPASy Home page