 ExPASy Home page ExPASy Home page |
Site Map | Search ExPASy | Contact us | Swiss-Prot | Proteomics tools |
 ExPASy Home page ExPASy Home page |
Site Map | Search ExPASy | Contact us | Swiss-Prot | Proteomics tools |
| Hosted by NCSC US | Mirror sites: | Canada | China | Korea | Switzerland | Taiwan |
An amino acid scale is defined by a numerical value assigned to each type of amino acid. The most frequently used scales are the hydrophobicity or hydrophilicity scales and the secondary structure conformational parameters scales, but many other scales exist which are based on different chemical and physical properties of the amino acids. This program provides 50 predefined scales entered from the literature.
You can set several parameters that control the computation of a scale profile, such as the window size, the weight variation model, the window edge relative weight value, and scale normalization.
The decrease in weight between the weight of the center and that of the edges will either be linear or exponential depending on the setting of the weight variation model option.
This option divides the weight into equally spaced intervals between 100% and the window edge relative weight (here: 10%).
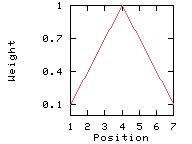
Weights used for the computation of the score for residue i: (window size 7, weight at window edges 10%) residue number i-3 i-2 i-1 i i+1 i+2 i+3 window position 1 2 3 4 5 6 7 --------------------------------------------------------------------------- weight 10% 40% 70% 100% 70% 40% 10%
This option makes the weights decrease exponentially from the central position to the window edge. This parameter has an effect only if you set the window edge relative weight to a value other than 100%.
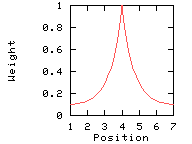
Weights used for the computation of the score for residue i: (window size 7, weight at window edges 10%) residue number i-3 i-2 i-1 i i+1 i+2 i+3 window position 1 2 3 4 5 6 7 --------------------------------------------------------------------------- weight 10% 12% 39% 100% 39% 12% 10%
 ExPASy Home page ExPASy Home page |
Site Map | Search ExPASy | Contact us | Swiss-Prot | Proteomics tools |
| Hosted by NCSC US | Mirror sites: | Canada | China | Korea | Switzerland | Taiwan |