http://www.blocks.fhcrc.org/
There is a tutorial on BLOCKS here: BLOCKS.
This database is made up of families described in InterPro. The clasification has some advantages because it allows us to easily see the short
conserved regions that are characteristic of a family.
The best thing about BLOCKS is that the web interface allows you to run multiple jobs, see the profiles you have constrcuted, construct trees, search other databases with the motifs you have constructed ...
Tour: we will search by "keyword", introducing "cytosine and methylase", thanks to which we will end up in IPB001525 (IPR001525 is the InterPro code). If we click here we will be able to see a series of motifs: IPB001525A, IPB001525B, IPB001525C, IPB001525D, IPB001525E and IPB001525F.
Clicking on Block Map will allow us to to see the distribution of these motifs in other proteins.
IPB001525: C-5 cytosine-specific DNA methylaseWe can also see the profiles that make up each motif graphically by clicking on "Logos". Or a tree of the sequences via ProWeb TreeViewer. Clicking on "Structures" we can see the 3D structures of these proteins (when they are available) and the distribution of the motifs around the structures.
6 distinct blocks in 158 sequences
MTA1_ARTLU|P31974 ( 521) -A-----BB-CCC---DDD--------------------------EE-F-----
MTB6_BACSP|P43420 ( 315) A-----BB-CCC--DDD----------EE-F--
MTB1_BACBR|P34905 ( 374) -A------BB-CCC--DDD-----------EE-F----
MTD2_HERAU|P25265 ( 354) -A-----BB-CCC--DDD------------EE--F---
MTA1_RUEGE|P94147 ( 429) A-----BB-CCC--DDD---------------------EE-F--
(...)
On top of that we can also use BLOCKS to search sequence databases via the following routes:
- COBBLER sequence: constructs a consensus sequences with the motif regions and searches the database with BLAST.
- MAST Search: generates PSSM (profiles) for each motif and searches with them. Here is an example.
- LAMA search: a tool for comparing multiple alignments useful fro finding homology between different families. An example:
- Search Pfam to see if the hits coincide with each other.
- Take a look at the "logos";
- Run BLAST; check the functional annotations;
- If both had 3D structures we could also compare these to see if they were similar.
- You can also check to see if the proteins that appear in one BLOCK also appear in the other.
- The tyrosine kinases, according to InterPro, can be defined with 3 "signatures" (markers):
- The serine/threonine kinases can be defined with two markers:
- LAMA version 28 Apr 00.
Minimal length of reported alignments 4
Score cutoff is 5.6 Z score units (in the top 7.7e-05 percentile
of chance scores)
alignment Z-score expected number for
block 1 from:to
block 2 from:to length
searching 5000 blocks
IPB001525A 1 : 14 and IPB001525A
1 : 14 (14) score 100 (19.2 0.0e+00)
IPB001525A 2 : 14 and IPB001566A
3 : 15 (13) score 47 ( 6.2 1.3e-01)
IPB001525A 1 : 13 and IPB001614A
28 : 40 (13) score 45 ( 5.7 3.5e-01)
IPB001525B 1 : 16 and IPB001525B
1 : 16 (16) score 100 (21.1 0.0e+00)
IPB001525B 2 : 16 and IPB001532D
11 : 25 (15) score 41 ( 5.6 3.7e-01)
IPB001525C 9 : 27 and IPB000146D
3 : 21 (19) score 37 ( 5.7 3.5e-01)
IPB001525C 2 : 23 and IPB000352A
26 : 47 (22) score 36 ( 5.6 3.6e-01)
IPB001525C 1 : 28 and IPB001525C
1 : 28 (28) score 100 (31.8 0.0e+00)
IPB001525C 8 : 28 and IPB002857C
1 : 21 (21) score 43 ( 7.2 1.3e-02)
IPB001525D 1 : 27 and IPB001525D
1 : 27 (27) score 100 (31.2 0.0e+00)
IPB001525D 1 : 27 and IPB004000C
15 : 41 (27) score 29 ( 5.9 2.0e-01)
IPB001525E 1 : 16 and IPB001525E
1 : 16 (16) score 100 (21.1 0.0e+00)
IPB001525E 3 : 16 and IPB002584A
6 : 19 (14) score 46 ( 6.0 1.7e-01)
IPB001525E 1 : 16 and IPB002857F
16 : 31 (16) score 65 (12.1 0.0e+00)
IPB001525F 1 : 10 and IPB001525F
1 : 10 (10) score 100 (14.5 0.0e+00)
IPB001525F 1 : 10 and IPB002190D
20 : 29 (10) score 57 ( 6.5 5.5e-02)
In fact in this case the two families are not related, its just that in defing the methylases a series of motifs were used that appear in various different domains. The methylase domain architecture is:
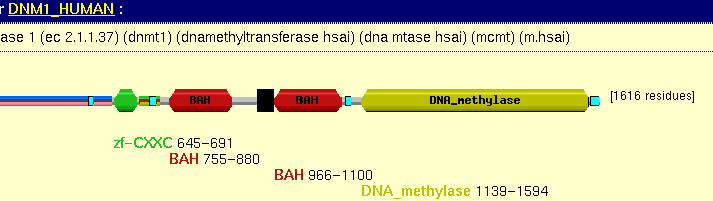
In fact there are many other families in BLOCKS that contain the RING-Zn zinc finger domain, so the two entries overlap.
(http://bioinf.man.ac.uk/dbbrowser/PRINTS/)
PRINTS Tutorial
The philosophy of this database is similar to that of BLOCKS.
It groups the proteins in superfamilies, families and subfamilies depending
on the presence of groups of tiny motifs - the digital "fingerprints" that characterise protein families.
A couple of examples of fingerprints: FASRECEPTOR and IL1BCENZYME.
You can do searches with
FPScan
to determine which fingerprints are similar to a determined sequence.
Here is an example, the results of a search with RASH_HUMAN.
(http://smart.embl-heidelberg.de/;
mirror en http://smart.ox.ac.uk/)
[ Smart Modular Architecture Research
Tool
]
The domains are constructed in a similar way to Pfam. Seed HMMs alignments are constructed for known domains
and the HMMs are used to search the databases for more proteins. For every domain they show the species
that are present, which other domains are associated with the domain, the 3D structure etc.
It is very similar to Pfam, although the definition of domains can vary between the two databases.
For exemple, according to Pfam there are 137
proteins that have the caspase domain, while in SMART there are
only 127.
[http://prodes.toulouse.inra.fr/prodom.html]
There is a tutorial at
this page.
Another database of domains. In the case of ProDom all the classifications are done automatially. using the program MKDOM2.
This database can be useful when a domain definition does not exist in the manually
curated databases like Pfam, Prosite or Smart.
[http://www.ebi.ac.uk/interpro/]
There is documentation in: http://www.ebi.ac.uk/interpro/user_manual.html?
As we can see there are a lot of databases for motifs and domains and each one has its own way of categorising proteins, its own distinct methods. InterPro unifies these databases under one single entry. InterPro is constructed from Pfam, SMART, Prosite, Prints, ProDom and TIGRFams.
| Database | Version | Entries |
| SWISS-PROT | 41.0 | 122564 |
| PRINTS | 35.0 | 1750 |
| TrEMBL | 23.0 | 830525 |
| Pfam | 8.0 | 5193 |
| PROSITE patterns | 17.37 | 1605 |
| PROSITE preprofiles | N/A | 150 |
| ProDom | 2002.1 | 1021 |
| InterPro | 6.0 | 7751 |
| Smart | 3.4 | 654 |
| TIGRFAMs | 2.1 | 1614 |
87% of the proteins in Swiss-Prot (and 75% of those in TrEMBL) have at least one connection with InterPro.
Example 1: the domain CARD. Along with the functional description of this domain, the page also shows that there are 117 proteins with this domain. It indicates that the corresponding domains in Pfam, Prosite and Smart (PF00619, PS50209 and SM00114) contain 82, 115 and 72 proteins, respectivly.
In the link "overview" (and in "...sorted by name" and in "detailed") they show graphically in which regions of each of the 117 the Pfam, Prosite and Smart domains appear. For each of the 117 proteins they also show which other domains are present. We can see that among these proteins there are several with the domain "protein kinase", others with the domain "caspase", etc. In "detailed" you can see the same thing in more detail. For example, in "overview" we can see this for protein RIK2_HUMAN:
![]()
We can see that the protein PIAP_PIG has various domains. The two at the start (in blue) are "BIR repeats" (the name comes up if you let the mouse hover over the domains), then there is a CARD domain and finally the RING Zn-finger domain.
In "detailed" we can see this:

You can see that in the case of the protein PIAP_PIG the domain CARD has three markers: Pfam, Prosite and Smart. The repeated BIR has 4 markers (PF0065, PS01282, PS50143 and SM00238), and the Zn-finger domains are found with 3 markers. Next to each marker we can see to which InterPro entry they correspond.
In the link "table", we can see that regions of each of the 117 proteins correspond with each of the "signatures".
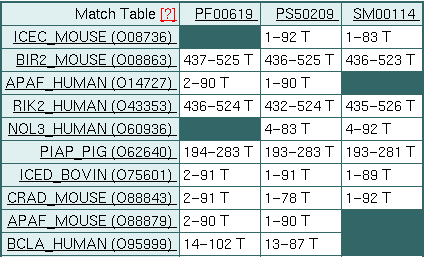
We can also see which of the "signatures" (markers) appear in each protein. In PIAP_PIG all three appear, but in NOL3_HUMAN, for example, the Pfam signature does not appear. It also indicates through the letters "T", "F" and "?", if the relationship is correct (T, true), incorrect (F, false) or uncertain (?).
What function could PIAP_PIG have? This protein is anotated in Swiss-Prot as a possible inhibitor of apoptosis. The CARD domain is a protein interaction domain. It normally interacts with other CARD domains, such as those present in caspases, kinases, etc. (the caspases are the principle executioners of apoptosis in the cell). At the same time the BIR domain (or BIR repeats) confers resistance to apoptosis. In addition, BIR normally appears associated with Zn-RING finger domain, which itself is implicated in protein-protein interactions.
We have rapidly gained some idea of the function of this protein from the database, and its possible role as an
inhibitor of apoptosis. It appears that it may interfere with the apoptotic machinery, perhaps via the interacting CARD and
zinc finger domains. This type of Zn-finger domain associated with many cellular processes, for example ubiquitination
(ticketing of proteins for destruction), where they interact with ubiquitin enzymes.
Hierarchical Classification in InterPro
Example 2: Protein kinases. Searching with the SwissProt sequence:
RAF1_HUMAN:
>RAF1_HUMAN|P04049|RAF proto-oncogene serine/threonine-protein kinase (EC 2.7.1.-) (RAF-1) (C-RAF).We can search via InterProScan (by clicking on "Sequence Search"). We have to give an email address, but any address will do.
MEHIQGAWKT ISNGFGFKDA VFDGSSCISP TIVQQFGYQR RASDDGKLTD PSKTSNTIRV
FLPNKQRTVV NVRNGMSLHD CLMKALKVRG LQPECCAVFR LLHEHKGKKA RLDWNTDAAS
LIGEELQVDF LDHVPLTTHN FARKTFLKLA FCDICQKFLL NGFRCQTCGY KFHEHCSTKV
PTMCVDWSNI RQLLLFPNST IGDSGVPALP SLTMRRMRES VSRMPVSSQH RYSTPHAFTF
NTSSPSSEGS LSQRQRSTST PNVHMVSTTL PVDSRMIEDA IRSHSESASP SALSSSPNNL
SPTGWSQPKT PVPAQRERAP VSGTQEKNKI RPRGQRDSSY YWEIEASEVM LSTRIGSGSF
GTVYKGKWHG DVAVKILKVV DPTPEQFQAF RNEVAVLRKT RHVNILLFMG YMTKDNLAIV
TQWCEGSSLY KHLHVQETKF QMFQLIDIAR QTAQGMDYLH AKNIIHRDMK SNNIFLHEGL
TVKIGDFGLA TVKSRWSGSQ QVEQPTGSVL WMAPEVIRMQ DNNPFSFQSD VYSYGIVLYE
LMTGELPYSH INNRDQIIFM VGRGYASPDL SKLYKNCPKA MKRLVADCVK KVKEERPLFP
QILSSIELLQ HSLPKINRSA SEPSLHRAAH TEDINACTLT TSPRLPVF
The result should be this.
We can see that it has an ras-interacting domain (domain RBD), un domain de union a diacil-glicerol (DAG_binding), and a protein kinase domain. If we look at the entry IPR000719, we can see that the family is the protein kinases and that it is subdivided into subfamilies, such as the serine/threonine kinases and the tyrosine kinases.

To which of the subfamilies does raf1_human belong?
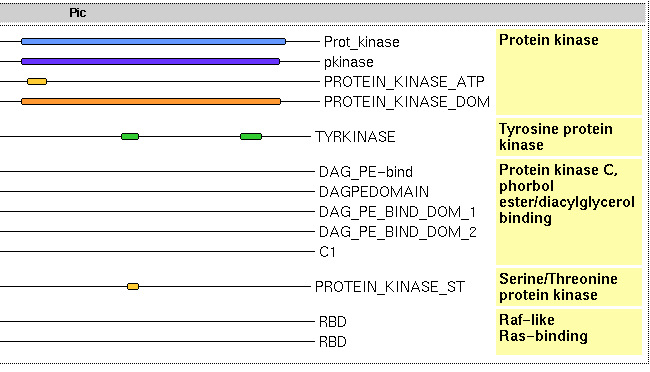
It is similar to both the tyrosine kinases and the serine/threonine kinases.
Which is correct?
Consult the entries IPR001245
y IPR002290.
PR00109;TYRKINASE (2753 proteins)
PS00109;PROTEIN_KINASE_TYR (1421 proteins)
SM00219;TyrKc (2090 proteins)
of which only that of PRINTS appears in raf1_human. This family is defined by 5 motifs in PRINTS, of which RAF1_HUMAN only has two.
PS00108;PROTEIN_KINASE_ST (6170 proteins)
SM00220;S_TKc
This last pattern from Prosite appears in the protein. The precision of this pattern is indicted in the Prosite entry:
[LIVMFYC]-x-[HY]-x-D-[LIVMFY]-K-x(2)-N-[LIVMFYCT](3)
Precision (true hits / (true hits + false positives)): 99.89 %
Recall (true hits / (true hits + false negatives)): 92.04 %
Conclusion: we would be inclined to conclude that the protein is a serine/threonine kinase,
given that the Prosite pattern has a high precision. [The annotation in Swiss-Prot -RAF1_HUMAN-
agrees with this hypothesis].
Practical Question: Is there any evolutionary relationship between the protein ras/p21 in humans and the elongation factor EF-Tu of E. coli?
Here are the sequences:
>RASH_HUMAN|P01112|Transforming protein P21/H-RAS-1 (C-H-RAS).
MTEYKLVVVG AGGVGKSALT IQLIQNHFVD EYDPTIEDSY RKQVVIDGET CLLDILDTAG
QEEYSAMRDQ YMRTGEGFLC VFAINNTKSF EDIHQYREQI KRVKDSDDVP MVLVGNKCDL
AARTVESRQA QDLARSYGIP YIETSAKTRQ GVEDAFYTLV REIRQHKLRK LNPPDESGPG
CMSCKCVLS
>EFTU_ECOLI|P02990|Elongation factor Tu (EF-Tu) (P-43).
SKEKFERTKP HVNVGTIGHV DHGKTTLTAA ITTVLAKTYG GAARAFDQID NAPEEKARGI
TINTSHVEYD TPTRHYAHVD CPGHADYVKN MITGAAQMDG AILVVAATDG PMPQTREHIL
LGRQVGVPYI IVFLNKCDMV DDEELLELVE MEVRELLSQY DFPGDDTPIV RGSALKALEG
DAEWEAKILE LAGFLDSYIP EPERAIDKPF LLPIEDVFSI SGRGTVVTGR VERGIIKVGE
EVEIVGIKET QKSTCTGVEM FRKLLDEGRA GENVGVLLRG IKREEIERGQ VLAKPGTIKP
HTKFESEVYI LSKDEGGRHT PFFKGYRPQF YFRTTDVTGT IELPEGVEMV MPGDNIKMVV
TLIHPIAMDD GLRFAIREGG RTVGAGVVAK VLS
If you want you can try BLAST. But the best local alignment you could get is small with a low significance:Is there another way to find out? What conclusions do you come to? Is it possible to detect an evolutionary relationship? Do they two protiens have related domains? If so, how significant is the match? What other things can you detect from the E-values?
ras : ESRQAQDLARSYGIPYI
eftu: QTREHILLGRQVGVPYI