There is no known structure with sequence similarity to our query.
In this case, swissprot indicates it is a hypothetical protein containing a Flavin reductase like domain.
- Some Meta PP server results: second and third predictions.
- FFAS03
- Genesilico Metaserver
FSSP comparison results file
Alignment structure-sequence : text format
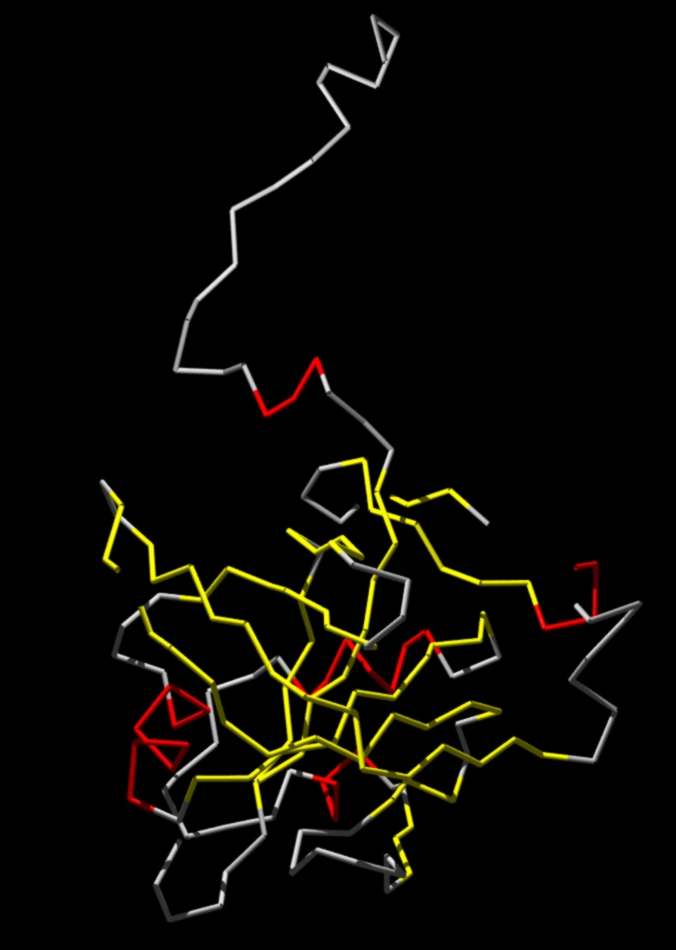
YELLOW: B-Strands.
RED: Helices.
GREY: No secondary structure elements.
Note the loops (are missing)
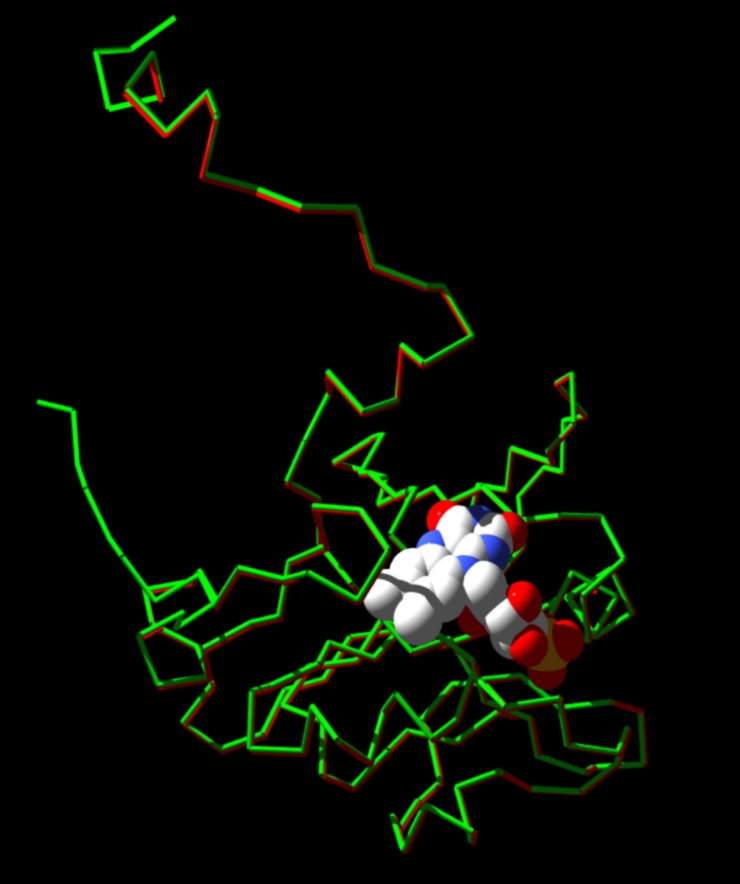
A comparison between the predicted structure and the actual one:
RED: The
predicted model.
GREEN: The template.
REST: FMN (flavin) group and Ni heteratms.
And, [HERE],the PDB file of the above comparison structure.