Practical Template-Based Modelling
includes
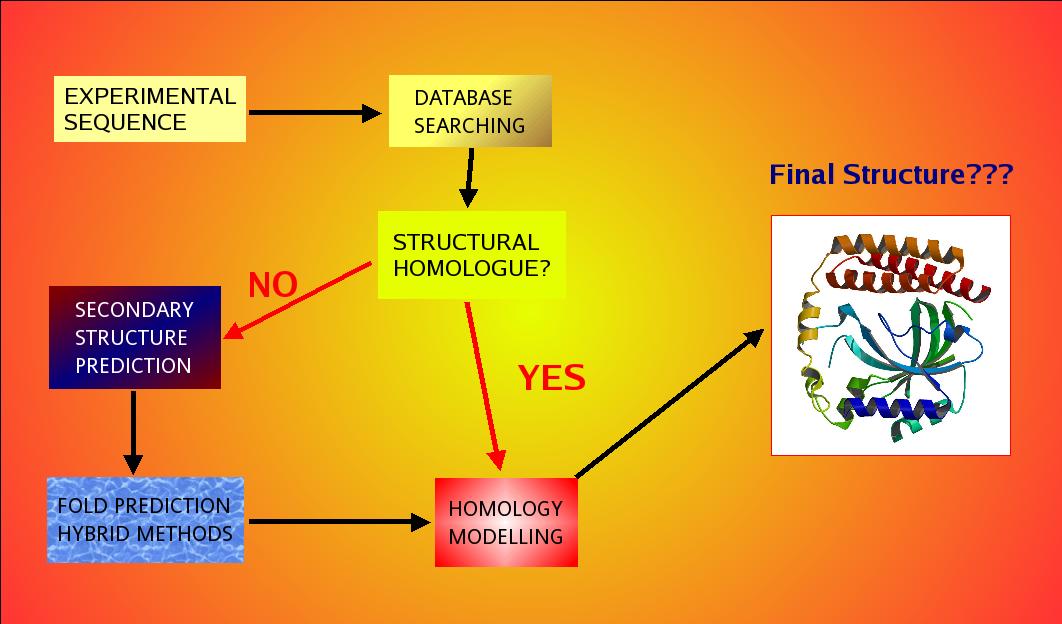
homology modelling, fold recognition and visualisation
 |
| This site is a complementary guide for the theroretical part of the course. You'll find examples and useful resources regarding protein structure prediction. This is a GUIDE, it is not intended to be a tutorial or similar resource. The practical parts have been divided in several blocks. |