
|
Swiss-Prot Protein Knowledgebase
Release Notes
Release 41, February 2003
|
1 Introduction
2 Description of the changes made to Swiss-Prot since release 40
3 Forthcoming changes
4 Status of the documentation files
5 The ExPASy World-Wide Web server
6 TrEMBL - a supplement to Swiss-Prot
7 FTP access to Swiss-Prot and TrEMBL
8 ENZYME and PROSITE
9 We need your help!
Appendix A
Appendix B
Release 41.0 of Swiss-Prot contains 122'564 sequence entries,
comprising 44'986'459 amino acids abstracted from 103'486
references. This represents an increase of 20% over release 40.0. The
growth of the database is summarized below.
-
| Release
| Date
| Number of entries
| Number of amino acids
|
| 2.0 |
09/86 |
3'939 |
900'163 |
| 3.0 |
11/86 |
4'160 |
969'641 |
| 4.0 |
04/87 |
4'387 |
1'036'010 |
| 5.0 |
09/87 |
5'205 |
1'327'683 |
| 6.0 |
01/88 |
6'102 |
1'653'982 |
| 7.0 |
04/88 |
6'821 |
1'885'771 |
| 8.0 |
08/88 |
7'724 |
2'224'465 |
| 9.0 |
11/88 |
8'702 |
2'498'140 |
| 10.0 |
03/89 |
10'008 |
2'952'613 |
| 11.0 |
07/89 |
10'856 |
3'265'966 |
| 12.0 |
10/89 |
12'305 |
3'797'482 |
| 13.0 |
01/90 |
13'837 |
4'347'336 |
| 14.0 |
04/90 |
15'409 |
4'914'264 |
| 15.0 |
08/90 |
16'941 |
5'486'399 |
| 16.0 |
11/90 |
18'364 |
5'986'949 |
| 17.0 |
02/91 |
20'024 |
6'524'504 |
| 18.0 |
05/91 |
20'772 |
6'792'034 |
| 19.0 |
08/91 |
21'795 |
7'173'785 |
| 20.0 |
11/91 |
22'654 |
7'500'130 |
| 21.0 |
03/92 |
23'742 |
7'866'596 |
| 22.0 |
05/92 |
25'044 |
8'375'696 |
| 23.0 |
08/92 |
26'706 |
9'011'391 |
| 24.0 |
12/92 |
28'154 |
9'545'427 |
| 25.0 |
04/93 |
29'955 |
10'214'020 |
| 26.0 |
07/93 |
31'808 |
10'875'091 |
| 27.0 |
10/93 |
33'329 |
11'484'420 |
| 28.0 |
02/94 |
36'000 |
12'496'420 |
| 29.0 |
06/94 |
38'303 |
13'464'008 |
| 30.0 |
10/94 |
40'292 |
14'147'368 |
| 31.0 |
02/95 |
43'470 |
15'335'248 |
| 32.0 |
11/95 |
49'340 |
17'385'503 |
| 33.0 |
02/96 |
52'205 |
18'531'384 |
| 34.0 |
10/96 |
59'021 |
21'210'389 |
| 35.0 |
11/97 |
69'113 |
25'083'768 |
| 36.0 |
07/98 |
74'019 |
26'840'295 |
| 37.0 |
12/98 |
77'977 |
28'268'293 |
| 38.0 |
07/99 |
80'000 |
29'085'965 |
| 39.0 |
05/00 |
86'593 |
31'411'114 |
| 40.0 |
10/01 |
101'602 |
37'315'215 |
| 41.0 |
02/03 |
122'564 |
44'986'459 |
|
2 Description of the changes made to Swiss-Prot
since release 40
|
-
2.1 Sequences and annotations
21'133 sequences have been added since release 40, the sequence data of
3'251 existing entries has been updated and the annotations of 57'525
entries have been revised.
-
2.2 The HPI project
The Human Proteomics Initiative (HPI) puts a major effort on the annotation
of all known human sequences according to the quality standards of Swiss-Prot.
This means that, for each known protein, a wealth of information is provided,
which includes the description of its function, its domain structure, subcellular
location, post-translational modifications (PTMs), variants, similarities to other
proteins, etc. This not only implies the annotation of newly detected proteins,
but also the integration of new research data into the existing entries by
specialized biologists, who are in close contact with experts all over the
world.
There are currently 9'172 annotated human sequences in Swiss-Prot.
Up-to-date detailed statistics concerning the HPI project are available
at:
Simultaneously, two further efforts were increased: the description of
human diseases associated with deficiency(ies) in the protein, and mammalian
orthologs of human proteins are annotated at a level equivalent to that of
the cognate human sequences.
For all aspects of the HPI project, we would appreciate the help and
collaboration of the scientific community. Information concerning the human
proteome is highly critical to a large section of the life science
community. We therefore appeal to the user community to fully participate
in this initiative by providing all the necessary information to define and
to speed up the comprehensive annotation of the human proteome.
For a detailed description of the HPI project please consult:
-
2.3 The HAMAP project
The first complete microbial genome sequence was that of the bacterium
Haemophilus influenzae, which became available in 1995. Since then, more
than 100 bacterial and archaeal genomes have been sequenced and many more
sequencing projects of pathogenic and nonpathogenic microbes are in
progress. To date, the publicly available microbial genomes
encode more than 230'000 different proteins.
In order to handle the large amount of "raw" data coming from microbial
genome sequencing, the High quality Automated Microbial Annotation of
Proteomes (HAMAP) project was initiated. The project aims to automatically
annotate a significant percentage of protein sequences, which originate from
microbial genome sequencing projects.
To maintain a high level quality of annotation, specific tools are
developed to deal with two completely separate subsets of bacterial and
archaeal proteins: proteins that have no recognizable similarity to any
other microbial or non-microbial proteins ("ORFans") and proteins that are
part of well-defined families or subfamilies. This is done by using a rule
system that describes the level and extent of annotations that can be
assigned by similarity with a prototype manually annotated entry. The
result is a curated entry whose quality is identical to that produced
manually by an expert annotator.
Programs under development are designed to recognize protein
peculiarities, and only proteins which match the defined criteria are
processed automatically. Protein sequences which fail to fit into the rule
system are further analyzed by Swiss-Prot expert annotators.
For a detailed description of the HAMAP project and its current status
please consult:
and:
Gattiker A., Michoud K., Rivoire C., Auchincloss A.H., Coudert E., Lima T., Kersey P.,
Pagni M., Sigrist C.J.A., Lachaize C., Veuthey A.-L., Gasteiger E., Bairoch A.
Automatic annotation of microbial proteomes in Swiss-Prot.
Comput. Biol. Chem. 27:49-58(2003).
-
2.4 What's happening with the model organisms?
We have selected a number of organisms that are the target of genome sequencing and/or
mapping projects and for which we intend to:
- be as complete as possible. All sequences available at a given time should
be immediately included in Swiss-Prot. This also includes sequence
corrections and updates;
- provide a higher level of annotation;
- provide cross-references to specialized database(s) that contain, among
other data, some information about the genes that code for these proteins;
- provide specific indexes and documents.
From our efforts to annotate human sequence entries as completely as possible
arose the HPI project (see 2.2), and the bacterial model organisms became
the focus of the HAMAP project (see 2.3). Here is the current status of the
model organisms which are not covered by these two projects:
-
-
2.5 'Nucleomorph' added to the OrGanelle (OG) line
The OG (OrGanelle) line indicates from which genome a gene for a protein
originates. Until now, defined terms in the OG line where 'Chloroplast',
'Cyanelle', 'Mitochondrion' and 'Plasmid'.
The term 'Nucleomorph' has been added, which is the
residual nucleus of an algal endosymbiont that resides inside its host cell.
-
2.6 Progress in the conversion of Swiss-Prot to mixed-case characters
We are gradually converting Swiss-Prot entries from all 'UPPER CASE'
to 'MiXeD CaSe'. With this release the RC (Reference Comment) line
topic STRAIN and the CC line topic 'CATALYTIC ACTIVITY' have been converted.
As described in section 3.2, the process of converting all of Swiss-Prot to mixed
case continues.
-
2.7 Multiple RP lines
Starting with release 41, there can be more than one RP (Reference
Position) line per reference in a Swiss-Prot entry. The RP line describes
the extent of the work carried out by the authors of the reference, e.g.
the type of molecule that has been sequenced, protein characterization,
PTM characterization, protein structure analysis, variation detection, etc.
As the number of experimental results per publication has increased over
the years, the limitation of using a single RP line per reference no longer
allowed to add all the information while maintaining a consistent format.
Therefore we decided to permit multiple RP lines.
Example:
RP SEQUENCE FROM N.A., SEQUENCE OF 23-42 AND 351-365, AND
RP CHARACTERIZATION.
-
2.8 Changes concerning cross-references (DR line)
-
2.8.1 Schizosaccharomyces pombe GeneDB database
We have added cross-references to the Schizosaccharomyces pombe GeneDB database (available at
http://www.genedb.org/genedb/pombe/index.jsp),
which contains all S. pombe known and predicted protein coding genes, pseudogenes and tRNAs.
It is hosted by the Sanger Institute.
The identifiers of the appropriate DR line are:
| Data bank identifier: |
GeneDB_SPombe |
| Primary identifier: |
GeneDB's unique identifier for a S. pombe gene. |
| Secondary identifier: |
None; a dash '-' is stored in that field. |
| Example: |
DR GeneDB_SPombe; SPAC9E9.12c; -. |
-
2.8.2 Genew
We have added cross-references to the Human Gene Nomenclature Database Genew (available at
http://www.gene.ucl.ac.uk/nomenclature/searchgenes.pl),
which provides data for all human genes which have approved symbols. It is managed
by the HUGO Gene Nomenclature Committee (HGNC).
The identifiers of the appropriate DR line are:
| Data bank identifier: |
Genew |
| Primary identifier: |
HGNC's unique identifier for a human gene |
| Secondary identifier: |
HGNC's approved gene symbol. |
| Example: |
DR Genew; HGNC:5217; HSD3B1. |
-
2.8.3 Gramene
We have added cross-references to the Gramene database, a comparative mapping resource for grains
(available at http://www.gramene.org/).
The format for the explicit links are:
| Data bank identifier: |
Gramene |
| Primary identifier: |
Unique identifier for a protein, which is identical to the
Swiss-Prot primary AC number of that protein. |
| Secondary identifier: |
None; a dash '-' is stored in that field. |
| Example: |
DR Gramene; Q06967; -. |
-
2.8.4 HAMAP
We have added cross-references to the collection of orthologous microbial protein families, generated manually
by expert curators of the HAMAP (High-quality Automated and Manual Annotation of microbial Proteomes) project
in the framework of the Swiss-Prot protein knowledgebase. The data is accessible at
http://www.expasy.org/sprot/hamap/families.html.
The identifiers of the appropriate DR line are:
| Data bank identifier: |
HAMAP |
| Primary identifier: |
HAMAP unique identifier for a microbial protein family |
| Secondary identifier: |
The values are either '-', 'fused', 'atypical' or 'atypical/fused'. The value '-' is a placeholder for an
empty field; the 'fused' value indicates that the family rule does not cover the entire protein; the value
'atypical' points out that the protein is divergent in sequence or has mutated functional sites, and should not be included in family
datasets. The value 'atypical/fused' indicates both latter findings. |
| Tertiary identifier: |
Number of domains found in the protein, generally '1', rarely '2' for the fusion of 2 identical domains. |
| Example: |
DR HAMAP; MF_00012; -; 1. |
-
2.8.5 Phosphorylation Site Database
We have added cross-references to the Phosphorylation Site Database, PhosSite (available at
http://vigen.biochem.vt.edu/xpd/xpd.htm),
which provides access to information from scientific literature concerning
prokaryotic proteins that undergo covalent phosphorylation on the hydroxyl side
chains of serine, threonine or tyrosine residues.
The identifiers of the appropriate DR line are:
| Data bank identifier: |
PhosSite |
| Primary identifier: |
Unique identifier for a phosphoprotein, which is identical to the
Swiss-Prot primary AC number of that protein.
|
| Secondary identifier: |
None; a dash '-' is stored in that field. |
| Example: |
DR PhosSite; P00955; -. |
-
2.8.6 TIGRFAMs
We have added cross-references to TIGRFAMs, a protein family database available at
http://www.tigr.org/TIGRFAMs/.
The identifiers of the appropriate DR line are:
| Data bank identifier: |
TIGRFAMs |
| Primary identifier: |
TIGRFAMs unique identifier for a protein family. |
| Secondary identifier: |
TIGRFAMs entry name for a protein family. |
| Tertiary identifier: |
Number of hits found in the sequence. |
| Example: |
DR TIGRFAMs; TIGR00630; uvra; 1. |
-
2.8.7 CarbBank
We have removed the Swiss-Prot cross-references to CarbBank.
-
2.8.8 GCRDb
We have removed the Swiss-Prot cross-references to GCRDb.
-
2.8.9 Mendel
We have removed the Swiss-Prot cross-references to Mendel.
-
2.8.10 YEPD
We have removed the Swiss-Prot cross-references to the yeast electrophoresis protein
database (YEPD).
-
2.9 Explicit links to dbSNP in FT VARIANT lines of human sequence entries
In human protein sequence entries we have introduced explicit links to the Single
Nucleotide Polymorphism database (dbSNP)
from the feature description of FT VARIANT keys.
The format of such links is:
FT VARIANT from to description (IN dbSNP:accession_number).
FT /FTId=VAR_number.
Example:
FT VARIANT 65 65 T -> I (IN dbSNP:1065419).
FT /FTId=VAR_012009.
-
2.10 Feature key 'SIMILAR' became obsolete
The feature key 'SIMILAR' was used to describe the extent of a similarity with another
protein sequence. Nowadays, most domains with similarity to other proteins are known
regions described in domain and family databases, which are annotated in Swiss-Prot with
the feature key 'DOMAIN' or 'REPEAT' and the comment (CC) line topic 'SIMILARITY'; thus the
feature key 'SIMILAR' became obsolete and will not be used again.
-
2.11 Version of SP in XML format
A distribution version of Swiss-Prot and TrEMBL in XML format is being
developed. The first draft of the XML specification was released for public
review on February 21, 2002.
For more information see http://www.ebi.ac.uk/swissprot/SP-ML/.
Please send comments and suggestions by electronic mail to sp-ml@ebi.ac.uk.
Please note that these are the last release notes in this format. In future,
forthcoming changes and recent modifications are announced to users also
between major Swiss-Prot releases.
The distinct sections of this document will move to the following sites:
|
-
3.1 Extension of the entry name format
We endeavor to assign meaningful entry names that facilitate the
identification of the proteins and the species of origin. Currently
the entry name consists of up to ten uppercase
alphanumeric characters. Swiss-Prot uses a general purpose naming
convention that can be symbolized as X_Y, where X is a mnemonic code of at
most 4 alphanumeric characters representing the protein name, the
'_' sign serves as a separator, and the Y is a mnemonic species
identification code of at most 5 alphanumeric characters representing the
biological source of the protein.
We are planning to elongate the mnemonic code for the protein name from up
to 4 characters to up to 5 characters. E.g. the mnemonic code for the
meiotic recombination protein rec10 is currently 'RE10'. After the
introduction of extended entry names it could be modified to the 5-letter
code 'REC10'.
-
3.2 Continuation of the conversion of Swiss-Prot to mixed-case characters
We will continue to convert Swiss-Prot entries from all 'UPPER CASE'
to 'MiXeD CaSe'. We are proceeding in the conversion of CC (Comment) lines,
we will start to convert the GN (Gene Name) lines to mixed case, but also any other
line type might be effected.
-
3.3 Reference Comment (RC) line topics may span lines
The RC (Reference Comment) line store comments relevant to the reference
cited, in currently 5 distinct topics: PLASMID, SPECIES, STRAIN, TISSUE and
TRANSPOSON. It is not always possible to list all information within one line.
Therefore we will allow multiple RC lines, in which one topic might span over
a line. Example:
RC STRAIN=Various strains;
could become
RC STRAIN=AZ.026, DC.005, GA.039, GA2181, IL.014, IN.018, KY.172, KY2.37,
RC LA.013, MN.001, MNb027, MS.040, NY.016, OH.036, TN.173, TN2.38,
RC UT.002, AL.012, AZ.180, MI.035, VA.015, and IL2.17;
-
3.4 New format of comment line (CC) topics
We are continuing a major overhaul of various comment line topics. We would
like the majority of the information stored to be usable by computer
programs (while remaining human-readable). We are therefore standardizing the
format of the topics.
-
3.4.1 ALTERNATIVE PRODUCTS
We are gradually restructuring the CC (comment) line topic ALTERNATIVE PRODUCTS
and introducing unique identifiers for each described isoform. Qualifiers,
which will be introduced are described in the table below:
-
| Topic |
Description |
| Event |
Biological process that results in the production of the alternative
forms (Alternative promoter, Alternative splicing, Alternative initiation).
Format: Event=controlled vocabulary;
Example: Event=Alternative splicing; |
| Named isoforms |
Number of isoforms listed in the topics 'Name' below the topic 'Event=Alternative splicing'.
Format: Named isoforms=number;
Example: Named isoforms=6; |
| Comment |
Any comments concerning one or more isoforms; optional; may be longer than 1 line.
Format: Comment=free text;
Example: Comment=Experimental confirmation may be lacking for some isoforms; |
| Name |
A common name for an isoform used in the literature or assigned by Swiss-Prot (currenty only available for spliced isoforms).
Format: Name=common name;
Example: Name=Alpha; |
Synonyms |
Synonyms for an isoform as used in the literature; optional.
Format: Synonyms=synonym_1[, synonym_n];
Example: Synonyms=B, KL5; |
| IsoId |
Unique identifier for an isoform, consisting of the Swiss-Prot accession
number, followed by a dash and an identifier for this isoform.
Format: IsoId=acc#-isoform_number[, acc#-isoform_number];
Example: IsoId=P05067-1; |
| Sequence |
Lists all FT VARSPLIC identifiers (VSP_#), which are needed to build the sequence
for a specific isoform. If the accession number of the IsoId does not correspond
to the accession number of the current entry, this topic contains the term
'External'.
Format: Sequence=VSP_#[, VSP_#]|Displayed|External|Not described;
Example: Sequence=Displayed;
Example: Sequence=VSP_000013, VSP_000014;
|
| Note |
Notes concerning current isoform; optional;
Format: Note=free text;
Example: Note=Predicted; |
In the case of 'Alternative initiation' the topic 'Event' can be
followed by a 'Comment' of free text. Format:
CC -!- ALTERNATIVE PRODUCTS:
CC Event=Alternative initiation;
CC Comment=Optional free text with information on alternative
CC initiation or the products retrieved from this event. In the
CC case of alternative initiation there will be no other topics;
In the case of 'Alternative splicing' the topic 'Event' can be followed by a
'Comment' of free text and a listing of all described isoforms. Format:
CC -!- ALTERNATIVE PRODUCTS:
CC Event=Alternative splicing;
CC Comment=Optional free text with information on alternative
CC splicing or the products retrieved from this event;
CC Name=isoform_1; Synonyms=synonym_1[, synonym_n];
CC IsoId=isoform_identifier_1[, isoform_identifer_n];
CC Sequence=VSP_identifier_1 [, VSP_identifier_n];
CC Note=Optional note concerning isoform_1;
CC Name=isoform_n; Synonyms=synonym_1[, synonym_n];
CC IsoId=isoform_identifier_1[, isoform_identifer_n];
CC Sequence=VSP_identifier_1 [, VSP_identifier_n];
CC Note=Optional note concerning isoform_n;
Example for new format of the CC lines and the corresponding FT lines for an entry
with alternative splicing:
...
CC -!- ALTERNATIVE PRODUCTS:
CC Event=Alternative splicing; Named isoforms=9;
CC Comment=Additional isoforms seem to exist. APP695, APP751 and
CC APP770 are the major isoforms. The L-isoforms are referred to as
CC appicans. Experimental confirmation may be lacking for some
CC isoforms;
CC Name=APP770; Synonyms=Prea4 770;
CC IsoId=P05067-1; Sequence=Displayed;
CC Name=APP305;
CC IsoId=P05067-2; Sequence=VSP_000005, VSP_000006;
CC Name=L-APP677;
CC IsoId=P05067-3; Sequence=VSP_000002, VSP_000004, VSP_000009;
CC Name=APP695; Synonyms=Prea4 695;
CC IsoId=P05067-4; Sequence=VSP_000002, VSP_000004;
CC Name=L-APP696;
CC IsoId=P05067-5; Sequence=VSP_000002, VSP_000003, VSP_000009;
CC Name=APP714;
CC IsoId=P05067-6; Sequence=VSP_000002, VSP_000003;
CC Name=L-APP733;
CC IsoId=P05067-7; Sequence=VSP_000007, VSP_000008, VSP_000009;
CC Name=APP751; Synonyms=Prea4 751;
CC IsoId=P05067-8; Sequence=VSP_000007, VSP_000008;
CC Name=L-APP752;
CC IsoId=P05067-9; Sequence=VSP_000009;
...
FT VARSPLIC 289 289 E -> V (in isoform APP695, isoform
FT L-APP696, isoform L-APP677 and isoform
FT APP714).
FT /FTId=VSP_000002.
FT VARSPLIC 290 345 Missing (in isoform L-APP696 and isoform
FT APP714).
FT /FTId=VSP_000003.
FT VARSPLIC 290 364 Missing (in isoform APP695 and isoform
FT L-APP677).
FT /FTId=VSP_000004.
FT VARSPLIC 290 305 VCSEQAETGPCRAMIS -> KWYKEVHSGQARWLML (in
FT isoform APP305).
FT /FTId=VSP_000005.
FT VARSPLIC 306 770 Missing (in isoform APP305).
FT /FTId=VSP_000006.
FT VARSPLIC 345 345 M -> I (in isoform L-APP733 and isoform
FT APP751).
FT /FTId=VSP_000007.
FT VARSPLIC 346 364 Missing (in isoform L-APP733 and isoform
FT APP751).
FT /FTId=VSP_000008.
FT VARSPLIC 637 654 Missing (in isoform L-APP677, isoform
FT L-APP696, isoform L-APP733 and isoform
FT L-APP752).
FT /FTId=VSP_000009.
...
-
3.4.2 PATHWAY
We are gradually structuring the comment line topic PATHWAY. To describe
the biochemical pathway in which the protein is involved, we use the
following format:
CC -!- PATHWAY: biochemical pathway; nth step.[ Comment.]
Example:
CC -!- PATHWAY: Coenzyme A (CoA) biosynthesis; first step.
-
3.4.3 COFACTOR
The comment line topic COFACTOR is gradually being modified to the following
format:
CC -!- COFACTOR: cofactor1[, cofactor2 and cofactor3].[ Comment.]
Examples:
CC -!- COFACTOR: Magnesium.
CC -!- COFACTOR: Copper, Manganese and Nickel.
-
3.5 Changes concerning cross-references (DR line)
We will add cross-references to the Gene Ontology (GO) database (available at
http://www.geneontology.org/), which
provides controlled vocabularies for the description of the molecular function,
biological process and cellular component of gene products.
The identifiers of the appropriate DR line are:
| Data bank identifier: |
GO |
| Primary identifier: |
GO's unique identifier for a GO term. |
| Secondary identifier: |
A 1-letter abbreviation for one of the 3 ontology aspects, separated from the GO term by a column. If the
term is longer than 45 characters, the first 43 characters are indicated followed by 3 dots ('...').
The abbreviations for the 3 distinct aspects of the ontology are P (biological Process), F (molecular Function) and
C (cellular Component). |
| Tertiary identifier: |
3-character GO evidence code.
|
| Example: |
DR GO; GO:0003677; F:DNA binding; TAS. |
-
3.6 Modifications concerning the feature table (FT line)
We are investigating a major effort in the annotation of posttranslational modifications, which has an
effect on various feature keys and feature descriptions. Major format changes are described below.
-
3.6.1 New feature key 'CROSSLNK'
The feature key 'CROSSLNK' will be introduced to describe bonds between amino
acids, which are formed posttranslationally within a peptide or between
peptides, such as isopeptidic bonds, carbon-carbon linkages, carbon-nitrogen
linkages and backbone condensations. It will also include the description of
tioether bonds and thiolester bonds and thus the feature keys 'THIOETH' and
'THIOLEST' will be removed.
Note: Disulfide bonds occur so often in proteins, that we will keep the
special feature key 'DISULFID' to describe this kind of linkage.
Format:
FT CROSSLNK from to Description.
-
3.6.2 Removal of the feature key 'THIOETH'
See section 3.6.1.
-
3.6.3 Removal of the feature key 'THIOLEST'
See section 3.6.1.
|
4 Status of the documentation files
|
Swiss-Prot is distributed with a large number of documentation files. Some
of these files have been available for a long time (the user manual,
release notes, the various indexes for authors, citations, keywords, etc.),
but many have been created recently and we are continuously adding new
files, and updating and modifying existing files. Please note that the
header in many documentation files has changed. The following table lists all
the documents that are currently available.
See also section 7.3 for information on how to access updated
versions of all documents between major releases.
| userman.txt |
User manual |
| relnotes.txt |
Release notes for the current release (41) |
| shortdes.txt |
Short description of entries in Swiss-Prot |
| |
|
| jourlist.txt |
List of cited journals |
| keywlist.txt |
List of keywords |
| plasmid.txt |
List of plasmids |
| speclist.txt |
List of organism (species) identification codes |
| tisslist.txt |
List of tissues |
| experts.txt |
List of on-line experts for PROSITE and Swiss-Prot |
| dbxref.txt |
List of databases cross-referenced in Swiss-Prot |
| submit.txt |
Submission of sequence data to Swiss-Prot |
| |
|
| acindex.txt |
Accession number index |
| autindex.txt |
Author index |
| citindex.txt |
Citation index |
| keyindex.txt |
Keyword index |
| speindex.txt |
Species index |
| deleteac.txt |
Deleted accession number index |
| |
|
| 7tmrlist.txt |
List of 7-transmembrane G-linked receptor entries |
| aatrnasy.txt |
List of aminoacyl-tRNA synthetases |
| allergen.txt |
Nomenclature and index of allergen sequences |
| annbioch.txt |
Swiss-Prot annotation: how is biochemical information assigned to sequence entries |
| arath.txt |
Index of Arabidopsis thaliana entries and their corresponding gene designations [see 2] |
| bacsu.txt |
Index of Bacillus subtilis strain 168 chromosomal entries and their
corresponding SubtiList cross-references [see 1] |
| bloodgrp.txt |
Blood group antigen proteins |
| bucai.txt |
Index of Buchnera aphidicola (subsp. Acyrthosiphon pisum) entries
[see 2] |
| bucap.txt |
Index of Buchnera aphidicola (subsp. Schizaphis graminum) entries[see 2] |
| calbican.txt |
Index of Candida albicans entries and their corresponding gene designations |
| cdlist.txt |
CD nomenclature for surface proteins of human leucocytes |
| celegans.txt |
Index of Caenorhabditis elegans entries and their corresponding gene
designations and WormPep cross-references |
| dicty.txt |
Index of Dictyostelium discoideum entries and their corresponding gene
designations and DictyDB cross-references |
| ec2dtosp.txt |
Index of Escherichia coli Gene-protein database (ECO2DBASE) entries
referenced in Swiss-Prot |
| ecoli.txt |
Index of Escherichia coli strain K12 chromosomal entries and their
corresponding EcoGene cross-references |
| embltosp.txt |
Index of EMBL Nucleotide Sequence Database entries referenced in Swiss-Prot |
| extradom.txt |
Nomenclature of extracellular domains |
| fly.txt |
Index of Drosophila entries and their corresponding FlyBase cross-references |
| glycosid.txt |
Classification of glycosyl hydrolase families and index of glycosyl hydrolase entries in Swiss-Prot |
| haein.txt |
Index of Haemophilus influenzae strain Rd chromosomal entries [see 1] |
| helpy.txt |
Index of Helicobacter pylori strain 26695 chromosomal entries [see 1] |
| hoxlist.txt |
Vertebrate homeotic Hox proteins: nomenclature and index |
| humchr01.txt |
Index of proteins encoded on human chromosome 1 |
| humchr02.txt |
Index of proteins encoded on human chromosome 2 |
| humchr03.txt |
Index of proteins encoded on human chromosome 3 |
| humchr04.txt |
Index of proteins encoded on human chromosome 4 |
| humchr05.txt |
Index of proteins encoded on human chromosome 5 |
| humchr06.txt |
Index of proteins encoded on human chromosome 6 |
| humchr07.txt |
Index of proteins encoded on human chromosome 7 |
| humchr08.txt |
Index of proteins encoded on human chromosome 8 |
| humchr09.txt |
Index of proteins encoded on human chromosome 9 |
| humchr10.txt |
Index of proteins encoded on human chromosome 10 |
| humchr11.txt |
Index of proteins encoded on human chromosome 11 |
| humchr12.txt |
Index of proteins encoded on human chromosome 12 |
| humchr13.txt |
Index of proteins encoded on human chromosome 13 |
| humchr14.txt |
Index of proteins encoded on human chromosome 14 |
| humchr15.txt |
Index of proteins encoded on human chromosome 15 |
| humchr16.txt |
Index of proteins encoded on human chromosome 16 |
| humchr17.txt |
Index of proteins encoded on human chromosome 17 |
| humchr18.txt |
Index of proteins encoded on human chromosome 18 |
| humchr19.txt |
Index of proteins encoded on human chromosome 19 |
| humchr20.txt |
Index of proteins encoded on human chromosome 20 |
| humchr21.txt |
Index of proteins encoded on human chromosome 21 |
| humchr22.txt |
Index of proteins encoded on human chromosome 22 |
| humchrx.txt |
Index of proteins encoded on human chromosome X |
| humchry.txt |
Index of proteins encoded on human chromosome Y |
| humpvar.txt |
Index of human proteins with sequence variants |
| initfact.txt |
List and index of translation initiation factors |
| intein.txt |
Index of intein-containing entries referenced in Swiss-Prot |
| metallo.txt |
Classification of metallothioneins and index of the entries in Swiss-Prot |
| metja.txt |
Index of Methanococcus jannaschii entries [see 1] |
| mgdtosp.txt |
Index of MGD entries referenced in Swiss-Prot |
| mimtosp.txt |
Index of MIM entries referenced in Swiss-Prot |
| mycge.txt |
Index of Mycoplasma genitalium strain G-37 chromosomal entries [see 1] |
| mycpn.txt |
Index of Mycoplasma pneumoniae strain M129 chromosomal entries [see 2] |
| ngr234.txt |
Table of predicted proteins in Rhizobium plasmid pNGR234a |
| nomlist.txt |
List of nomenclature related references for proteins |
| pdbtosp.txt |
Index of Protein Data Bank (PDB) entries referenced in Swiss-Prot |
| peptidas.txt |
Classification of peptidase families and index of peptidase entries in Swiss-Prot |
| plastid.txt |
List of chloroplast and cyanelle encoded proteins |
| pombe.txt |
Index of Schizosaccharomyces pombe entries and their corresponding gene designations |
| restric.txt |
List of restriction enzyme and methylase entries |
| ribosomp.txt |
Index of ribosomal proteins classified by families on the basis of sequence similarities |
| ricpr.txt |
Index of Rickettsia prowazekii strain Madrid E entries [see 1] |
| salty.txt |
Index of Salmonella typhimurium strain LT2 chromosomal entries and their
corresponding StyGene cross-references |
| syny3.txt |
Index of Synechocystis sp. strain PCC 6803 entries [see 1] |
| upflist.txt |
List of UPF (Uncharacterized Protein Families) and index of members |
| yeast.txt |
Index of Saccharomyces cerevisiae entries in Swiss-Prot and their
corresponding gene designations |
| yeast1.txt |
Yeast chromosome I entries |
| yeast2.txt |
Yeast chromosome II entries |
| yeast3.txt |
Yeast chromosome III entries |
| yeast5.txt |
Yeast chromosome V entries |
| yeast6.txt |
Yeast chromosome VI entries |
| yeast7.txt |
Yeast chromosome VII entries |
| yeast8.txt |
Yeast chromosome VIII entries |
| yeast9.txt |
Yeast chromosome IX entries |
| yeast10.txt |
Yeast chromosome X entries |
| yeast11.txt |
Yeast chromosome XI entries |
| yeast13.txt |
Yeast chromosome XIII entries |
| yeast14.txt |
Yeast chromosome XIV entries |
Notes:
| 1 |
The filenames for indexes of microbe-specific entries have been renamed; the filename
is now composed of the 5-letter code used for the species in the Swiss-Prot entry name
and the extension 'txt'. This modification concerns the following files:
'bacsu.txt' (formerly: 'subtilis.txt'), 'haein.txt' (formerly: 'haeinflu.txt'),
'helpy.txt' (formerly: 'hpylori.txt'), 'metja.txt' (formerly: 'mjannasc.txt'),
'mycge.txt' (formerly: 'mgenital.txt'), 'ricpr.txt' (formerly: 'rprowaze.txt'),
'syny3.txt' (formerly: 'pcc6803.txt').
|
| 2 |
The files 'arath.txt', 'bucai.txt', 'bucap.txt' and 'mycpn.txt' are new documents introduced since
release 40. |
We have continued to include in some Swiss-Prot documentation files the
references to Web sites relevant to the subject under consideration. There
are now 89 documents that include such links.
|
5 New features of the ExPASy World-Wide Web
server related to Swiss-Prot
|
Explicit general and continuously updated documentation about the ExPASy
server is available at
http://www.expasy.org/doc/expasy.pdf.
ExPASy is constantly modified and improved. If you wish to be informed on
the changes made to the server you can either:
- Read the document 'History of changes, improvements and new features' which is
available at the address: http://www.expasy.org/history.html
- Subscribe to Swiss-Flash, a service that reports news of databases, software and service
developments. By subscribing to this service, you will automatically get Swiss-Flash
bulletins by electronic mail. To subscribe, use the address:
http://www.expasy.org/swiss-flash/.
Among all the improvements and the new features introduced since the last
Swiss-Prot release, here are those that we believe are specifically useful
to Swiss-Prot users:
- The NiceProt view of Swiss-Prot has been further improved: access to documentation
has been facilitated by adding "mouse-over" hypertext links from various
sections in NiceProt to the corresponding information in the user manual.
Those hypertext links, which give access to documentation rather than the
data related to the protein entry, are visually different from the ordinary
hyperlinks. While they are not immediately recognizable as such, the user
can see that they
are clickable by moving the mouse pointer over the section headings such as
"References"
or
"Keywords".
A short description of the linked information appears at the bottom
of the web browser, and when clicked, a small additional window is opened with
related information extracted from the user manual.
Similarly, in the "Cross-references" section, the names of the databases
to which an entry is cross-referenced are linked to the corresponding sections
in the document dbxref.txt
(List of databases cross-referenced in Swiss-Prot).
- Implicit links have been added to the resources AraC-XylS, Ensembl and ModBase.
We have removed the implicit links to DOMO, which is no longer maintained.
For more details on Swiss-Prot cross-references, implicit and explicit links, you can read:
Gasteiger E., Jung E., Bairoch A.
Swiss-Prot: connecting biological knowledge via a protein database.
Curr. Issues Mol. Biol. 3:47-55(2001)
- A few improvements have been applied to the pages describing the
Human Proteomics Initiative (HPI).
For each human chromosome a link is provided to the corresponding index of
Swiss-Prot entries, to relevant information in the EBI Proteome database,
in Ensembl, in the Human Genome Resources at NCBI and in euGenes at Indiana
University.
The HPI status report
has been modified to include, for each of the counted items (e.g.
splice variants, variants, references) not only the absolute number,
but also the maximal and average number of occurrences per entry, and the number
of entries concerned by the counted item.
|
6 TrEMBL - a supplement to Swiss-Prot
|
The ongoing genome sequencing and mapping projects have dramatically
increased the number of protein sequences to be incorporated into Swiss-Prot.
Since we do not want to dilute the quality standards of Swiss-Prot by
incorporating sequences into the database without proper sequence analysis
and annotation, we cannot speed up the incorporation of new incoming data
indefinitely. But as we also want to make the sequences available as quickly
as possible, we introduced in 1995 a computer annotated supplement to
Swiss-Prot. This supplement consists of entries in Swiss-Prot-like format
derived from the translation of all coding sequences (CDS) in the EMBL
nucleotide sequence database, except those already included in Swiss-Prot.
This supplement is named TrEMBL (Translation from EMBL). It can be
considered as a preliminary section of Swiss-Prot. This Swiss-Prot release
is supplemented by TrEMBL release 21.
TrEMBL is available by FTP from the EBI
and ExPASy servers in the directory
'/databases/trembl'. It can be queried on WWW by the EBI and ExPASy
SRS servers. It is distributed with its own set of release notes.
|
7 FTP access to Swiss-Prot and TrEMBL
|
-
7.1 Generalities
Swiss-Prot is available for download on the following anonymous FTP
servers:
-
7.2 Non-redundant database
On the ExPASy and EBI FTP servers we distribute files that make up a non-redundant
and complete protein sequence database consisting of three components:
1) Swiss-Prot
2) TrEMBL
3) New entries to be integrated later into TrEMBL (hereafter known as TrEMBL_New)
Every week three files are completely rebuilt. These files are named:
sprot.dat.gz, trembl.dat.gz and trembl_new.dat.gz. As indicated by their '.
gz' extension, these are gzip-compressed files which, when
decompressed, produce ASCII files in Swiss-Prot format.
Three other files are also available (sprot.fas.gz, trembl.fas.gz and
trembl_new.fas.gz) which are compressed 'fasta' format sequence
files useful for building the databases used by FASTA, BLAST and other
sequence similarity search programs. Please do not use these files for any
other purpose, as you will lose all annotations by using this stripped-down
format.
The files for the non-redundant database are stored in the directory
'/databases/sp_tr_nrdb' on the ExPASy FTP server (ftp.expasy.org) and in
the directory '/pub/databases/sp_tr_nrdb' on the EBI FTP server
(ftp.ebi.ac.uk).
Additional notes:
- The Swiss-Prot file continuously grows as new annotated sequences are added.
- The TrEMBL file decreases in size as sequences are moved out of that section after being
annotated and moved into Swiss-Prot. Four times a year a new release of TrEMBL is
built at EBI, at this point the TrEMBL file increases in size as it then includes all of the new
data (see next section) that has accumulated since the last release.
- The TrEMBL_New file starts as a very small file and grows in size until a new release of
TrEMBL is available.
- Swiss-Prot and TrEMBL share the same system of accession numbers. Therefore you
will not find any primary accession number duplicated between the two sections. A TrEMBL
entry (and its associated accession number(s)) can either move to Swiss-Prot as a new
entry or be merged with an existing Swiss-Prot entry. In the latter case, the accession
number(s) of that TrEMBL entry are added to that of the Swiss-Prot entry.
- TrEMBL_New does not have real accession numbers. However it was necessary to have an
'AC' line so as to be able to use it with different software products. This AC line contains a
temporary identifier which consists of the protein_ID (protein sequence identifier) of the
coding sequence in the parent nucleotide sequence.
- TrEMBL_New is quite messy! You will of course find new sequence entries but you will also
encounter sequences that are going to be used to update existing TrEMBL or Swiss-Prot
entries. None of the "cleaning" steps that are applied to produce a TrEMBL release are run
on TrEMBL_New nor are any of the computer-annotation software tools that are used to
enhance the information content of TrEMBL. TrEMBL_New is provided only so that users
can be sure not to miss any important new sequences when they run similarity searches.
- While these three files allow you to build what we call a 'non-redundant' database, it must be
noted that this is not completely a true statement. Without going into a long explanation we
can say that this is currently the best attempt in providing a complete selection of protein
sequence entries while trying to eliminate redundancies. While Swiss-Prot is completely
(well 99.994% !) non-redundant, TrEMBL is far from being non-redundant and the addition of
Swiss-Prot + TrEMBL is even less so.
- To describe to your users the version of the non-redundant database that you are providing
them with, you should use a statement of the form:
Swiss-Prot release 41.x of xx-yyy-2003;
TrEMBL release 23.x of xx-yyy-2003;
TrEMBL_New of xx-yyy-2003.
-
7.3 Weekly updates of Swiss-Prot documents
Whilst the ExPASy FTP server so far only allowed FTP access to the
Swiss-Prot documents and indexes in their versions at the time of the
last full release, all documents are now updated with every weekly
release of Swiss-Prot. They are available for FTP download from the
directory /databases/swiss-prot/updated_doc/.
-
7.4 Weekly updates of Swiss-Prot
Weekly updates of Swiss-Prot are available by anonymous FTP. Three files
are generated at each update:
| new_seq.dat |
Contains all the new entries since the last full release; |
| upd_seq.dat |
Contains the entries for which the sequence data has been updated since the last release; |
| upd_ann.dat |
Contains the entries for which one or more annotation fields have been updated since the last release. |
Important notes
- Although we try to follow a regular schedule, we do not promise to update these files every
week. In most cases two weeks may elapse between two updates.
- Instead of using the above files, you can, every week, download an updated copy of the
Swiss-Prot database. This file is available in the directory containing the non-redundant
database (see section 7.2).
-
8.1 The ENZYME nomenclature database
Release 30.0 of the ENZYME nomenclature database is distributed with
release 41 of Swiss-Prot. ENZYME release 30.0 contains information relative
to 4'136 enzymes. In this release, we have added a significant number of new
entries and we also updated many entries.
-
8.2 The PROSITE database
PROSITE now comes with its own release notes.
We welcome feedback from our users. We would especially appreciate your notifying
us if you find that sequences belonging to your field of expertise
are missing from the database. We also would like to be notified about
annotations to be updated, if, for example, the function of a protein has
been clarified or if new information about post-translational modifications
has become available. To facilitate this feedback we offer, on the ExPASy
WWW server, a form that allows the submission of updates and/or corrections
to Swiss-Prot:
It is also possible, from any entry in Swiss-Prot displayed by the ExPASy
server, to submit updates and/or corrections for that particular entry.
Finally, you can also send your comments by electronic mail to the
address:
Note that all update requests are assigned a unique
identifier of the form UR-Xnnnn (example: UR-A0123). This identifier is
used internally by the Swiss-Prot staff at SIB and EBI to track requests
and is also used in e-mail exchanges with the persons who have
submitted a request.
|
APPENDIX A: Some statistics
|
-
A.1 Amino acid composition
-
A.1.1 Composition in percent for the complete database
Ala (A) 7.72 Gln (Q) 3.92 Leu (L) 9.56 Ser (S) 6.98
Arg (R) 5.24 Glu (E) 6.54 Lys (K) 5.96 Thr (T) 5.51
Asn (N) 4.28 Gly (G) 6.90 Met (M) 2.36 Trp (W) 1.18
Asp (D) 5.27 His (H) 2.26 Phe (F) 4.06 Tyr (Y) 3.13
Cys (C) 1.60 Ile (I) 5.88 Pro (P) 4.88 Val (V) 6.66
Asx (B) 0.000 Glx (Z) 0.000 Xaa (X) 0.01
-
A.1.2 Classification of the amino acids by their frequency
Leu, Ala, Ser, Gly, Val, Glu, Lys, Ile, Thr, Asp, Arg, Pro, Asn, Phe,
Gln, Tyr, Met, His, Cys, Trp
-
A.2 Taxonomic origin
Total number of species represented in this release of Swiss-Prot: 7'778
The first twenty species represent 51'656 sequences: 42.1% of the total
number of entries.
-
A.2.1 Table of the frequency of occurrence of species
Species represented 1x: 3679
2x: 1206
3x: 619
4x: 403
5x: 273
6x: 251
7x: 192
8x: 146
9x: 120
10x: 66
11- 20x: 331
21- 50x: 250
51-100x: 84
>100x: 158
-
A.2.2 Table of the most represented species
------ --------- --------------------------------------------
Number Frequency Species
------ --------- --------------------------------------------
1 9172 Homo sapiens (Human)
2 6169 Mus musculus (Mouse)
3 4892 Saccharomyces cerevisiae (Baker's yeast)
4 4832 Escherichia coli
5 3442 Rattus norvegicus (Rat)
6 2402 Bacillus subtilis
7 2291 Caenorhabditis elegans
8 2116 Schizosaccharomyces pombe (Fission yeast)
9 1952 Arabidopsis thaliana (Mouse-ear cress)
10 1773 Haemophilus influenzae
11 1764 Drosophila melanogaster (Fruit fly)
12 1529 Methanococcus jannaschii
13 1485 Escherichia coli O157:H7
14 1389 Bos taurus (Bovine)
15 1371 Mycobacterium tuberculosis
16 1240 Salmonella typhimurium
17 1062 Gallus gallus (Chicken)
18 942 Shigella flexneri
19 919 Synechocystis sp. (strain PCC 6803)
20 914 Escherichia coli O6
21 876 Archaeoglobus fulgidus
22 839 Pseudomonas aeruginosa
23 838 Xenopus laevis (African clawed frog)
24 822 Sus scrofa (Pig)
25 771 Salmonella typhi
26 716 Aquifex aeolicus
27 704 Oryctolagus cuniculus (Rabbit)
28 687 Mycoplasma pneumoniae
29 670 Rhizobium meliloti (Sinorhizobium meliloti)
30 609 Vibrio cholerae
31 599 Treponema pallidum
32 581 Mycobacterium leprae
33 572 Buchnera aphidicola (subsp. Acyrthosiphon pisum)
34 560 Buchnera aphidicola (subsp. Schizaphis graminum)
35 536 Helicobacter pylori (Campylobacter pylori)
36 535 Rickettsia prowazekii
37 524 Yersinia pestis
38 519 Helicobacter pylori J99 (Campylobacter pylori J99)
39 519 Streptomyces coelicolor
40 494 Bacillus halodurans
41 491 Zea mays (Maize)
42 491 Methanobacterium thermoautotrophicum
43 486 Mycoplasma genitalium
44 480 Pasteurella multocida
45 454 Anabaena sp. (strain PCC 7120)
46 432 Lactococcus lactis (subsp. lactis) (Streptococcus lactis)
47 419 Thermotoga maritima
48 416 Oryza sativa (Rice)
49 405 Borrelia burgdorferi (Lyme disease spirochete)
50 404 Chlamydia trachomatis
51 403 Rhizobium sp. (strain NGR234)
52 393 Canis familiaris (Dog)
53 391 Chlamydia pneumoniae (Chlamydophila pneumoniae)
54 390 Neisseria meningitidis (serogroup B)
55 386 Neisseria meningitidis (serogroup A)
56 381 Chlamydia muridarum
57 366 Caulobacter crescentus
58 365 Pyrococcus horikoshii
59 359 Listeria monocytogenes
60 359 Clostridium acetobutylicum
61 357 Pyrococcus abyssi
62 354 Ralstonia solanacearum (Pseudomonas solanacearum)
63 352 Listeria innocua
64 352 Rhizobium loti (Mesorhizobium loti)
65 350 Streptococcus pneumoniae
66 346 Agrobacterium tumefaciens (strain C58 / ATCC 33970)
67 341 Nicotiana tabacum (Common tobacco)
68 337 Xylella fastidiosa
69 335 Deinococcus radiodurans
70 332 Ovis aries (Sheep)
71 326 Xanthomonas campestris (pv. campestris)
72 325 Halobacterium sp. (strain NRC-1)
73 320 Staphylococcus aureus (strain N315)
74 320 Campylobacter jejuni
75 317 Staphylococcus aureus (strain Mu50 / ATCC 700699)
76 316 Dictyostelium discoideum (Slime mold)
77 311 Clostridium perfringens
78 299 Sulfolobus solfataricus
79 297 Staphylococcus aureus (strain MW2)
80 290 Corynebacterium glutamicum (Brevibacterium flavum)
81 288 Pisum sativum (Garden pea)
82 287 Xanthomonas axonopodis (pv. citri)
83 285 Streptococcus pyogenes
84 283 Aeropyrum pernix
85 278 Pyrococcus furiosus
86 278 Staphylococcus aureus
87 269 Brucella melitensis
88 268 Bacteriophage T4
89 266 Neurospora crassa
90 265 Triticum aestivum (Wheat)
91 264 Candida albicans (Yeast)
92 261 Rickettsia conorii
93 258 Hordeum vulgare (Barley)
94 254 Vaccinia virus (strain Copenhagen)
95 251 Glycine max (Soybean)
96 250 Lycopersicon esculentum (Tomato)
97 248 Rhodobacter capsulatus (Rhodopseudomonas capsulata)
98 247 Thermoanaerobacter tengcongensis
99 246 Solanum tuberosum (Potato)
100 244 Pseudomonas putida
-
A.2.3 Taxonomic distribution of the sequences
Kingdom Sequences (% of the database)
Archaea 7119 ( 6%)
Bacteria 46344 ( 38%)
Eukaryota 60623 ( 49%)
Viruses 8478 ( 7%)
Within Eukaryota:
Category sequences (% of Eukaryota) (% of the complete database)
Human 9172 ( 15%) ( 7%)
Other Mammalia 16041 ( 26%) ( 13%)
Other Vertebrata 5806 ( 10%) ( 5%)
Viridiplantae 9581 ( 16%) ( 8%)
Fungi 9337 ( 15%) ( 8%)
Insecta 3352 ( 6%) ( 3%)
Nematoda 2504 ( 4%) ( 2%)
Other 4830 ( 8%) ( 4%)
-
A.3 Sequence size
-
A.3.1 Repartition of the sequences by size (excluding fragments)
From To Number From To Number
1- 50 2283 1001-1100 1127
51- 100 8420 1101-1200 796
101- 150 12542 1201-1300 550
151- 200 11267 1301-1400 379
201- 250 11387 1401-1500 305
251- 300 10019 1501-1600 213
301- 350 10039 1601-1700 166
351- 400 9804 1701-1800 118
401- 450 7435 1801-1900 128
451- 500 6547 1901-2000 106
501- 550 5067 2001-2100 59
551- 600 3400 2101-2200 96
601- 650 2753 2201-2300 99
651- 700 2015 2301-2400 57
701- 750 1766 2401-2500 56
751- 800 1474 >2500 326
801- 850 1101
851- 900 1142
901- 950 817
951-1000 704
-
A.3.2 Longest and shortest sequences
The shortest sequence is GRWM_HUMAN (P24272) : 3 amino acids.
The longest sequence is NEBU_HUMAN (P20929) : 6669 amino acids.
-
A.4 Journal citations
Note: the following citation statistics reflect the number of distinct
journal citations.
Total number of journals cited in this release of Swiss-Prot: 1'316
-
A.4.1 Table of the frequency of journal citations
Journals cited 1x: 496
2x: 167
3x: 84
4x: 61
5x: 46
6x: 47
7x: 26
8x: 25
9x: 22
10x: 11
11- 20x: 98
21- 50x: 98
51-100x: 39
>100x: 96
-
A.4.2 List of the most cited journals in Swiss-Prot
Nb Citations Journal name
-- --------- -------------------------------------------------------------
1 9138 Journal of Biological Chemistry
2 5013 Proceedings of the National Academy of Sciences of the U.S.A.
3 3631 Nucleic Acids Research
4 3612 Journal of Bacteriology
5 3381 Gene
6 2663 FEBS Letters
7 2598 Biochemical and Biophysical Research Communications
8 2429 European Journal of Biochemistry
9 2383 Biochemistry
10 2171 The EMBO Journal
11 2045 Nature
12 2024 Biochimica et Biophysica Acta
13 1821 Journal of Molecular Biology
14 1752 Genomics
15 1579 Cell
16 1542 Molecular and Cellular Biology
17 1243 Biochemical Journal
18 1146 Science
19 1123 Plant Molecular Biology
20 1117 Molecular and General Genetics
21 1068 Molecular Microbiology
22 855 Journal of Biochemistry
23 830 Virology
24 748 Human Molecular Genetics
25 693 Journal of Cell Biology
26 645 Nature Genetics
27 597 Journal of Virology
28 588 Plant Physiology
29 582 Human Mutation
30 579 Genes and Development
31 550 Oncogene
32 538 The American Journal of Human Genetics
33 530 Infection and Immunity
34 529 Yeast
35 516 Journal of Immunology
36 494 Journal of General Virology
37 469 Archives of Biochemistry and Biophysics
38 454 Structure
39 446 FEMS Microbiology Letters
40 433 Microbiology
41 394 Development
42 379 Human Genetics
43 376 Current Genetics
44 376 Nature Structural Biology
45 347 Genetics
46 343 Molecular and Biochemical Parasitology
47 335 Blood
48 317 Applied and Environmental Microbiology
49 313 Journal of Clinical Investigation
50 299 Molecular Endocrinology
51 283 DNA and Cell Biology
52 282 Protein Science
53 281 Journal of Molecular Evolution
54 276 Developmental Biology
55 276 Mammalian Genome
56 271 Biological Chemistry Hoppe-Seyler
57 251 Cancer Research
58 248 Journal of Experimental Medicine
59 246 Neuron
60 241 Immunogenetics
61 240 Mechanisms of Development
62 229 Journal of General Microbiology
63 228 Endocrinology
64 221 DNA Sequence
65 217 Acta Crystallographica, Section D
66 213 Hoppe-Seyler's Zeitschrift fur Physiologische Chemie
67 209 Molecular Biology of the Cell
68 207 The Plant Cell
69 203 Journal of Cell Science
70 191 Molecular Biology and Evolution
71 190 Brain Research. Molecular Brain Research
72 187 The Plant Journal
73 183 Journal of Neurochemistry
74 180 Journal of Neuroscience
75 160 Comparative Biochemistry and Physiology
76 158 Cytogenetics and Cell Genetics
77 156 DNA
78 154 Bioscience, Biotechnology, and Biochemistry
79 152 The Journal of Clinical Endocrinology and Metabolism
80 145 Toxicon
81 144 Molecular Pharmacology
82 143 Antimicrobial Agents and Chemotherapy
83 140 American Journal of Physiology
84 131 Biochimie
85 127 Bioorganicheskaia Khimiia
86 125 Virus Research
87 125 Proteins
88 122 DNA Research
89 121 Molecular Plant-Microbe Interactions
90 119 Hemoglobin
91 116 Peptides
92 114 Agricultural and Biological Chemistry
93 112 Current Biology
94 111 Journal of Investigative Dermatology
95 110 Molecular and Cellular Endocrinology
96 106 Genome Research
-
A.5 Statistics for some line types
The following table summarizes the total number of some Swiss-Prot lines,
as well as the number of entries with at least one such line, and the
frequency of the lines.
Total Number of Average
Line type / subtype number entries per entry
--------------------------------- -------- --------- ---------
References (RL) 232571 1.90
Journal 195556 111991 1.60
Submitted to EMBL/GenBank/DDBJ 34500 27873 0.28
Unpublished observations 536 532 <0.01
Submitted to Swiss-Prot 464 462 <0.01
Plant Gene Register 463 453 <0.01
Book citation 460 450 <0.01
Thesis 190 188 <0.01
Submitted to other databases 190 189 <0.01
Unpublished results 123 121 <0.01
Patent 87 86 <0.01
Worm Breeder's Gazette 2 2 <0.01
Comments (CC) 405433 3.31
SIMILARITY 117866 103489 0.96
FUNCTION 77092 75796 0.63
SUBCELLULAR LOCATION 55038 55038 0.45
CATALYTIC ACTIVITY 39528 37138 0.32
SUBUNIT 33846 33846 0.28
PATHWAY 17449 16966 0.14
TISSUE SPECIFICITY 13626 13626 0.11
COFACTOR 12141 12141 0.10
MISCELLANEOUS 7816 7190 0.06
PTM 7140 6571 0.06
ALTERNATIVE PRODUCTS 3946 3946 0.03
INDUCTION 3558 3558 0.03
DOMAIN 3535 3241 0.03
DEVELOPMENTAL STAGE 3362 3362 0.03
CAUTION 3342 3172 0.03
DISEASE 2244 1868 0.02
ENZYME REGULATION 1753 1753 0.01
MASS SPECTROMETRY 893 810 0.01
DATABASE 818 751 0.01
POLYMORPHISM 343 334 <0.01
BIOTECHNOLOGY 50 50 <0.01
PHARMACEUTICAL 47 47 <0.01
Features (FT) 655938 5.35
DOMAIN 95401 28727 0.78
TRANSMEM 77067 16988 0.63
CONFLICT 47337 16661 0.39
CARBOHYD 45507 11138 0.37
DISULFID 41846 10872 0.34
TURN 39177 2956 0.32
METAL 36827 10004 0.30
STRAND 36304 2644 0.30
HELIX 27742 2845 0.23
ACT_SITE 24322 15216 0.20
CHAIN 23456 19176 0.19
VARIANT 23307 4423 0.19
REPEAT 22336 3704 0.18
NP_BIND 15500 10893 0.13
SIGNAL 14828 14826 0.12
MOD_RES 13336 7528 0.11
NON_TER 10321 7875 0.08
BINDING 8145 6285 0.07
ZN_FING 7821 2770 0.06
VARSPLIC 6951 3249 0.06
SITE 6265 4319 0.05
INIT_MET 5574 5545 0.05
PROPEP 4686 4026 0.04
MUTAGEN 4273 1337 0.03
DNA_BIND 4193 3949 0.03
CA_BIND 4049 1149 0.03
LIPID 2946 2395 0.02
TRANSIT 2582 2562 0.02
PEPTIDE 2517 1001 0.02
NON_CONS 804 411 0.01
UNSURE 290 123 <0.01
SE_CYS 111 73 <0.01
THIOETH 94 32 <0.01
THIOLEST 23 23 <0.01
Cross-references (DR) 999237 8.15
EMBL 230657 116257 1.88
InterPro 195677 104236 1.60
Pfam 133012 99557 1.09
PROSITE 105218 66696 0.86
PIR 47040 35736 0.38
PRINTS 39413 34822 0.32
SMART 38729 29473 0.32
HSSP 38069 38069 0.31
TIGRFAMs 31394 29063 0.26
ProDom 30120 28820 0.25
HAMAP 23868 23778 0.19
PDB 11737 3547 0.10
TIGR 11065 11020 0.09
MIM 8171 7086 0.07
Genew 7836 7788 0.06
MGD 5820 5805 0.05
SGD 4936 4882 0.04
EcoGene 4228 4226 0.03
MEROPS 3316 3222 0.03
TRANSFAC 2464 2214 0.02
WormPep 2413 2239 0.02
SubtiList 2362 2361 0.02
FlyBase 2236 2173 0.02
GeneDB_SPombe 2131 2101 0.02
TubercuList 1400 1363 0.01
StyGene 1196 1193 0.01
SWISS-2DPAGE 810 809 0.01
ListiList 712 658 0.01
Leproma 585 581 <0.01
Gramene 411 411 <0.01
MaizeDB 405 401 <0.01
HIV 370 354 <0.01
REBASE 358 353 <0.01
ECO2DBASE 351 299 <0.01
DictyDb 319 316 <0.01
GlycoSuiteDB 259 259 <0.01
ZFIN 225 225 <0.01
PHCI-2DPAGE 211 211 <0.01
MypuList 131 131 <0.01
Aarhus/Ghent-2DPAGE 128 98 <0.01
Siena-2DPAGE 104 104 <0.01
HSC-2DPAGE 85 85 <0.01
PhosSite 53 53 <0.01
COMPLUYEAST-2DPAGE 50 50 <0.01
PMMA-2DPAGE 47 47 <0.01
Maize-2DPAGE 39 39 <0.01
SagaList 25 25 <0.01
ANU-2DPAGE 15 15 <0.01
-
A.6 Miscellaneous statistics
Total number of distinct authors cited in Swiss-Prot: 164'410
Total number of chloroplast-encoded sequences: 3'131
Total number of mitochondrial-encoded sequences: 2'385
Total number of cyanelle-encoded sequences: 145
Total number of plasmid-encoded sequences: 2'624
Number of additional sequences encoded in splice variants : 5'661
|
APPENDIX B: Relationships between Swiss-Prot and some biomolecular databases
|
The current status of the relationships (cross-references) between Swiss-Prot and some
biomolecular databases is shown in the following schema:
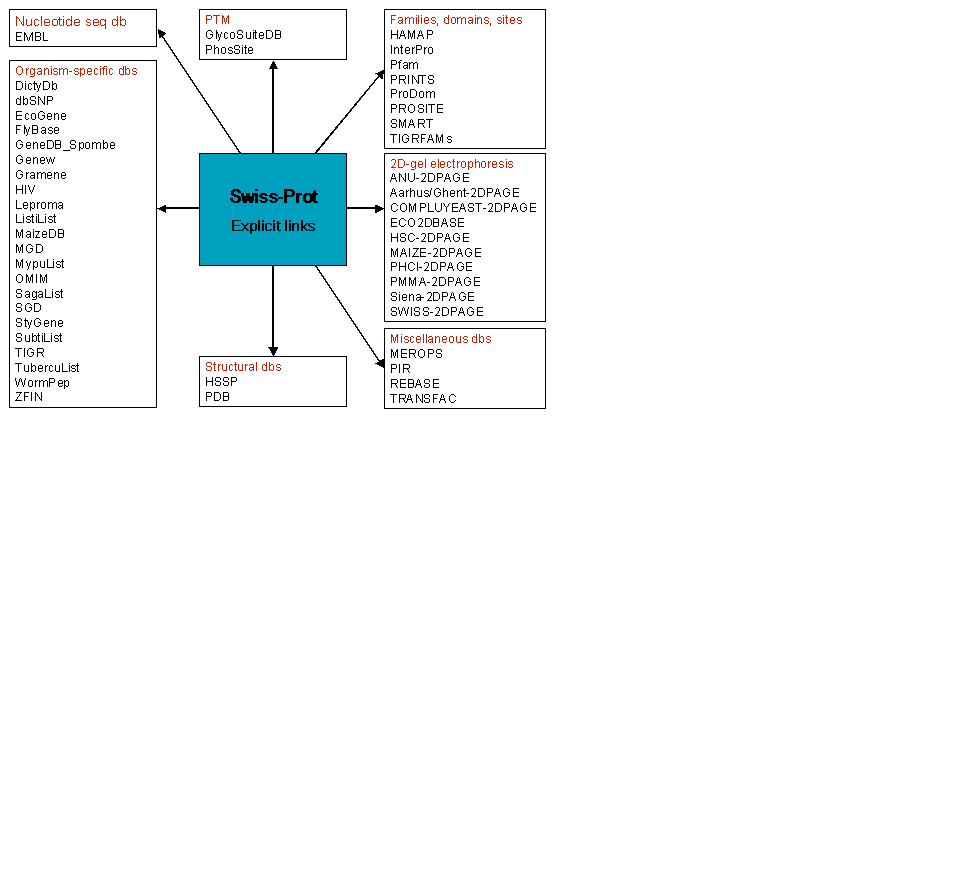
 ExPASy Home page
ExPASy Home page ExPASy Home page
ExPASy Home page

 ExPASy Home page
ExPASy Home page